MMBioS Research Highlights
2021 2020 2019 2018 2017 2016 2015 2014 2013
2021 Highlights
Direct coupling of oligomerization and oligomerization-driven endocytosis of the dopamine transporter to its conformational mechanics and activity (Sorkina T, Cheng MH, Bagalkot TR, Wallace C, Watkins SC, Bahar I, Sorkin A. J Biol Chem. 2021)
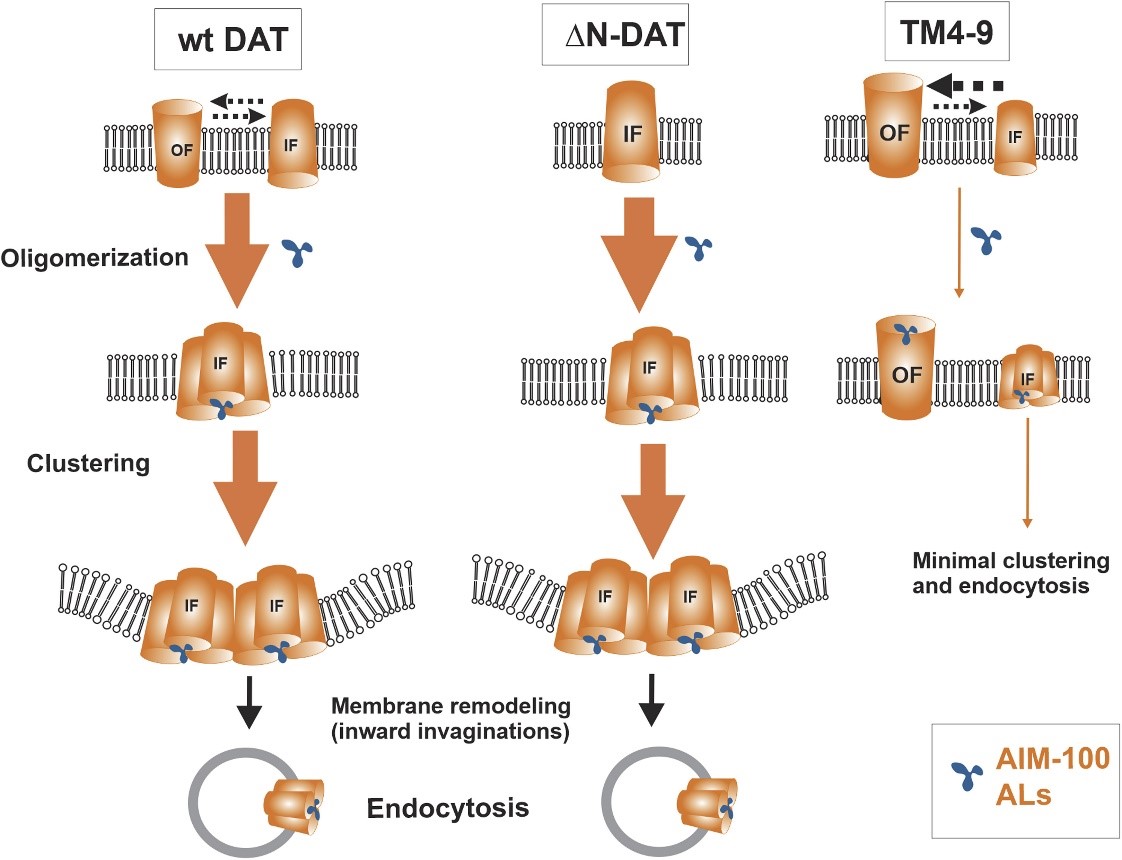
Dopamine transporter (DAT) mediates the reuptake of synaptically-released dopamine, and thus controls the duration and intensity of dopamine neurotransmission. Mammalian DAT has been observed to form oligomers, although the mechanisms of oligomerization and its role in DAT activity and trafficking remain largely unknown. We discovered a series of small molecule compounds that stabilize trimers and induce high-order oligomers of DAT, and concomitantly promote its clathrin-independent endocytosis. Using a combination of chemical cross-linking, FRET microscopy, antibody-uptake endocytosis assay, live-cell lattice light sheet microscopy, ligand binding and substrate transport kinetics analyses, and molecular modeling and simulations, we investigated molecular basis of DAT oligomerization and endocytosis induced by these compounds. Our study showed that small molecule-induced DAT oligomerization and endocytosis are favored by the inward-facing DAT conformation and involve interactions of four hydrophobic residues at the interface between transmembrane helices TM4 and TM9. Surprisingly, a corresponding quadruple DAT mutant displays altered dopamine transport kinetics and increased cocaine-analog binding. The latter is shown to originate from an increased preference for outward-facing conformation and inward-to-outward transition. Taken together, our results demonstrate a direct coupling between conformational dynamics of DAT, functional activity of the transporter and its oligomerization leading to endocytosis. The high specificity of such coupling for DAT makes the TM4-9 hub a new target for pharmacological modulation of DAT activity and subcellular localization.
|
Hypothetical model of the effects of AIM-100 and ALs on DAT. AIM-100 and ALs stabilize trimers (oligomers), promote formation of high-order oligomers and nanoclusters of a fraction of wt DAT in the IF conformation and ΔN-DAT, that is predominantly in the IF state. Clustering of concave-shaped DATs leads to membrane remodeling and stabilizing inward invaginations of the membrane, followed up by vesicle scission (endocytosis). The same ALs and AIM-100 are less efficient in inducing oligomerization of DAT in the OF conformation. For example, the capacity of AIM-100 and ALs to promote oligomerization of TM4-9 is diminished because of the increased probability of the OF state of this mutant. AIM-100 and ALs, are, however, still capable of binding at or near the substrate binding sites of the TM4-9 mutant in its OF state. Widths of arrows roughly reflect process probabilities. |
Sorkina T, Cheng MH, Bagalkot TR, Wallace C, Watkins SC, Bahar I, Sorkin A. (2021) Direct coupling of oligomerization and oligomerization-driven endocytosis of the dopamine transporter to its conformational mechanics and activity. J Biol Chem. Feb 18:100430. doi: 10.1016/j.jbc.2021.100430 PMID: 33610553
2020 Highlights
Differences in the intrinsic spatial dynamics of the chromatin contribute to cell differentiation (Zhang S, Chen F, Bahar I. Nucleic Acids Res. 2020)
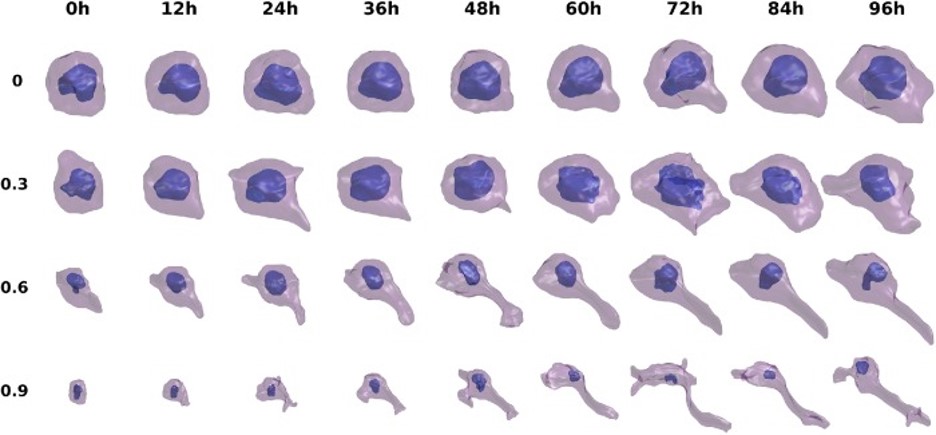
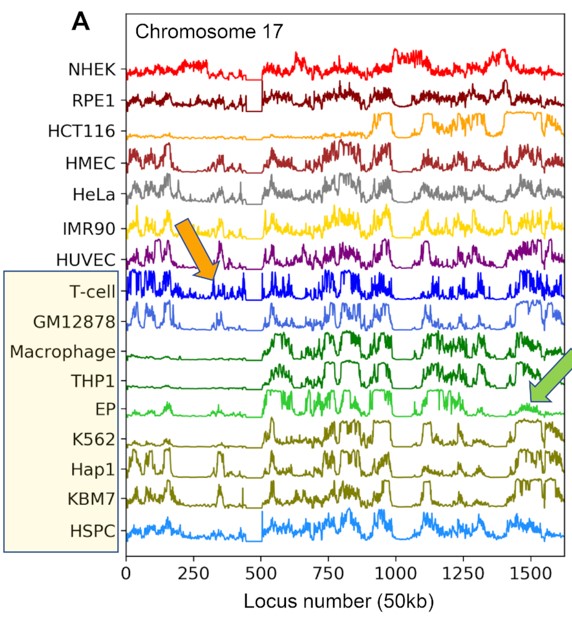 Advances in chromosome conformation capture techniques as well as computational characterization of genomic loci structural dynamics open new opportunities for exploring the mechanistic aspects of genome-scale differences across different cell types. We examined here the dynamic basis of variabilities between different cell types by investigating their chromatin mobility profiles inferred from Hi-C data using an elastic network model representation of the chromatin within the scope of our DBP6. Our comparative analysis of sixteen cell lines reveals close similarities between chromosomal dynamics across different cell lines on a global scale, but notable cell-specific variations emerge in the detailed spatial mobilities of genomic loci. Closer examination reveals that the differences in spatial dynamics mainly originate from the difference in the frequencies of their intrinsically accessible modes of motion. Thus, even though the chromosomes of different types of cells have access to similar modes of collective movements, not all modes are deployed by all cells, such that the effective mobilities and cross-correlations of genomic loci are cell-type-specific. Comparison with RNA-seq expression data reveals a strong overlap between highly expressed genes and those distinguished by high mobilities in the present study, in support of the role of the intrinsic spatial dynamics of chromatin as a determinant of cell differentiation.
Advances in chromosome conformation capture techniques as well as computational characterization of genomic loci structural dynamics open new opportunities for exploring the mechanistic aspects of genome-scale differences across different cell types. We examined here the dynamic basis of variabilities between different cell types by investigating their chromatin mobility profiles inferred from Hi-C data using an elastic network model representation of the chromatin within the scope of our DBP6. Our comparative analysis of sixteen cell lines reveals close similarities between chromosomal dynamics across different cell lines on a global scale, but notable cell-specific variations emerge in the detailed spatial mobilities of genomic loci. Closer examination reveals that the differences in spatial dynamics mainly originate from the difference in the frequencies of their intrinsically accessible modes of motion. Thus, even though the chromosomes of different types of cells have access to similar modes of collective movements, not all modes are deployed by all cells, such that the effective mobilities and cross-correlations of genomic loci are cell-type-specific. Comparison with RNA-seq expression data reveals a strong overlap between highly expressed genes and those distinguished by high mobilities in the present study, in support of the role of the intrinsic spatial dynamics of chromatin as a determinant of cell differentiation.
Zhang S, Chen F, Bahar I. Differences in the intrinsic spatial dynamics of the chromatin contribute to cell differentiation. Nucleic Acids Res 48: 1131-1145
Nanoscale co-organization and coactivation of AMPAR, NMDAR, and mGluR at excitatory synapses (Goncalves J, Bartol T, Camus C, Levet F, Menegolla AP, Sejnowski T, Sibarita JB, Vivaudou M, Choquet D, and Hosy E. Proc Natl Acad Sci USA. 2020)
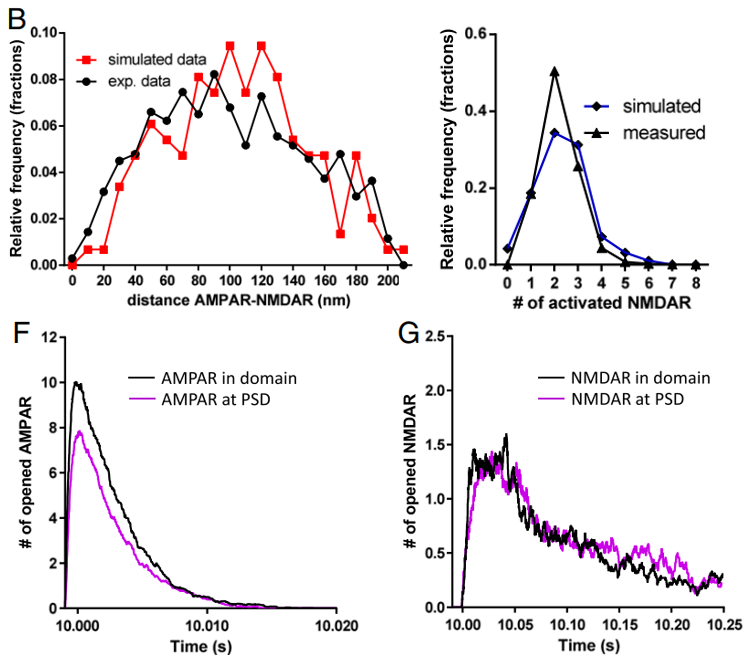 Neuron-to-neuron communication is based on synapse activity where presynapses release neurotransmitters, which activate postsynaptic neurotransmitter receptors. The various families of glutamate receptors at the excitatory synapses are responsible of the fast synaptic transmission as well as the regulation of the long-term signaling implicated in information storage and memory. The organization at the nanometer scale of these postsynaptic receptors is a key determinant for synaptic transmission efficiency. Here, we combined dual-color superresolution imaging with electrophysiology and modeling to determine how the various glutamate receptors are co-organized at the nanoscale and to what extent this organization regulates the receptor activation by a single vesicle release.
Neuron-to-neuron communication is based on synapse activity where presynapses release neurotransmitters, which activate postsynaptic neurotransmitter receptors. The various families of glutamate receptors at the excitatory synapses are responsible of the fast synaptic transmission as well as the regulation of the long-term signaling implicated in information storage and memory. The organization at the nanometer scale of these postsynaptic receptors is a key determinant for synaptic transmission efficiency. Here, we combined dual-color superresolution imaging with electrophysiology and modeling to determine how the various glutamate receptors are co-organized at the nanoscale and to what extent this organization regulates the receptor activation by a single vesicle release.
Goncalves J, Bartol T, Camus C, Levet F, Menegolla AP, Sejnowski T, Sibarita JB, Vivaudou M, Choquet D, and Hosy E. Nanoscale co-organization and coactivation of AMPAR, NMDAR, and mGluR at excitatory synapses. Proc Natl Acad Sci USA 117 (25) 14503-14511.
Parallel Tempering with Lasso for model reduction in systems biology (Gupta S, Lee REC, Faeder JR. PLOS Computational Biology. 2020)
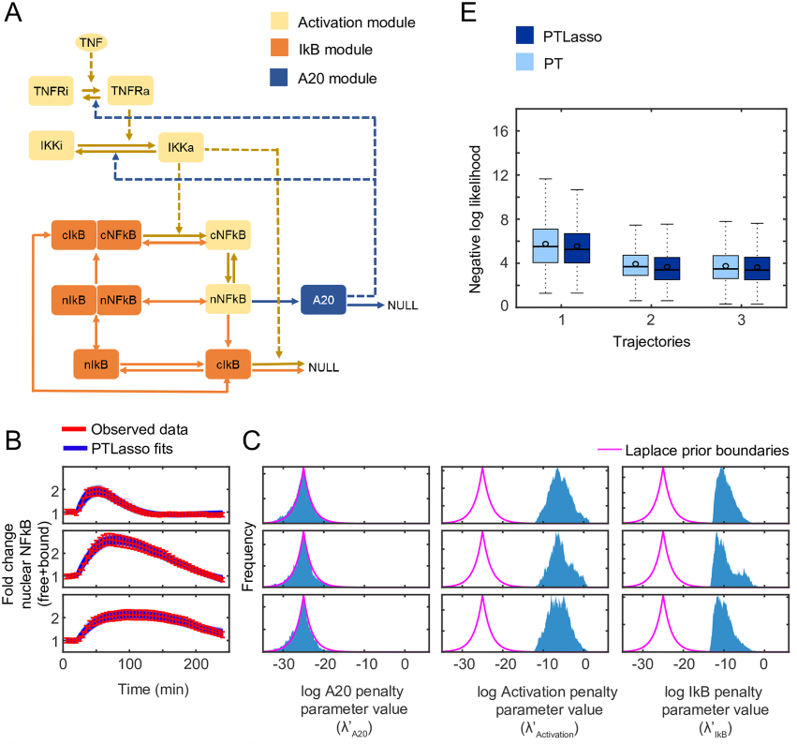 Here we demonstrate that parallel tempering (PT), a technique widely used in the molecular dynamics community to accelerate sampling of conformational space, could be used to improve the performance of Monte Carlo methods used in Bayesian parameter estimation. We have recently combined this approach with Lasso, a commonly-used regularization method in the machine learning field, to identify and reduce complex models of biochemical kinetics. Applying this method, PTLasso, as shown in the accompanying figure, by placing regularization constraints on different components of a signaling network, we can determine which elements of the network are required to produce a particular set of responses observed in experiments.
Here we demonstrate that parallel tempering (PT), a technique widely used in the molecular dynamics community to accelerate sampling of conformational space, could be used to improve the performance of Monte Carlo methods used in Bayesian parameter estimation. We have recently combined this approach with Lasso, a commonly-used regularization method in the machine learning field, to identify and reduce complex models of biochemical kinetics. Applying this method, PTLasso, as shown in the accompanying figure, by placing regularization constraints on different components of a signaling network, we can determine which elements of the network are required to produce a particular set of responses observed in experiments.
Gupta S, Lee REC, Faeder JR. Parallel Tempering with Lasso for model reduction in systems biology. PLOS Comput. Biol., 16(3), p. e1007669, 2020.
Image-derived models of cell organization changes during differentiation and drug treatments (Ruan X., et al. Molecular Biology of the Cell. 2020)
Our work on modeling PC12 cells undergoing differentiation into neuron-like morphologies (under C&SP11, completed) has been published in Molecular Biology of the Cell. We have also made the large dataset of 3D images collected in that study available through Dryad (https://doi.org/10.5061/dryad.hc8037v).
Abstract: PC12 cells are a popular model system to study changes driving and accompanying neuronal differentiation. While attention has been paid to changes in transcriptional regulation and protein signaling, much less is known about the changes in organization that accompany PC12 differentiation. Fluorescence microscopy can provide extensive information about these changes, although it is difficult to continuously observe changes over many days of differentiation. We describe a generative model of differentiation-associated changes in cell and nuclear shape and their relationship to mitochondrial distribution constructed from images of different cells at discrete time points. We show that the model accurately represents complex cell and nuclear shapes and learn a regression model that relates cell and nuclear shape to mitochondrial distribution; the predictive accuracy of the model increases during differentiation. Most importantly, we propose a method, based on cell matching and interpolation, to produce realistic simulations of the dynamics of cell differentiation from only static images. We also found that the distribution of cell shapes is hollow: most shapes are very different from the average shape. Finally, we show how the method can be used to model nuclear shape changes of human-induced pluripotent stem cells resulting from drug treatments.
Ruan X, Johnson GR, Bierschenk I, Nitschke R, Boerries M, Busch H, Murphy RF. Image-derived models of cell organization changes during differentiation and drug treatments. Mol Biol Cell, 2020. 31(7): p. 655-666.
2019 Highlights
Monoamine transporters: structure, intrinsic dynamics and allosteric regulation (Cheng and Bahar. Nature Structural & Molecular Biology. 2019)
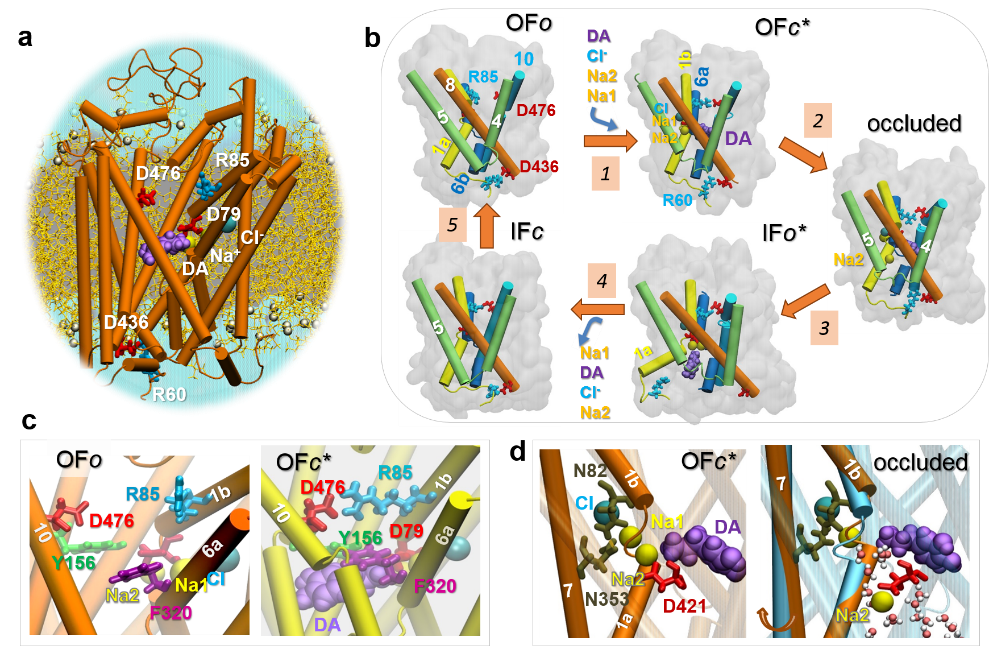
Monoamine transporters (MATs) regulate neurotransmission via the reuptake of dopamine, serotonin and norepinephrine from extra-neuronal regions and thus maintain neurotransmitter homeostasis. As targets of a wide range of compounds, including antidepressants, substances of abuse and drugs for neuropsychiatric and neurodegenerative disorders, their mechanism of action and their modulation by small molecules have long been of broad interest. Recent advances in the structural characterization of dopamine and serotonin transporters have opened the way for structure-based modeling and simulations, which, together with experimental data, now provide mechanistic understanding of their transport function and interactions. Here we review recent progress in the elucidation of the structural dynamics of MATs and their conformational landscape and transitions, as well as allosteric regulation mechanisms.
Cheng MH & Bahar I. (2019) Monoamine transporters: structure, intrinsic dynamics and allosteric regulation, Nature Structural & Molecular Biology | VOL 26 | JULY 2019 | 545–556
Trimerization of dopamine transporter triggered by AIM-100 binding (Cheng, Ponzoni, Sorkina, Lee, Zhang, Sorkin, Bahar. Neuropharmacology. 2019)
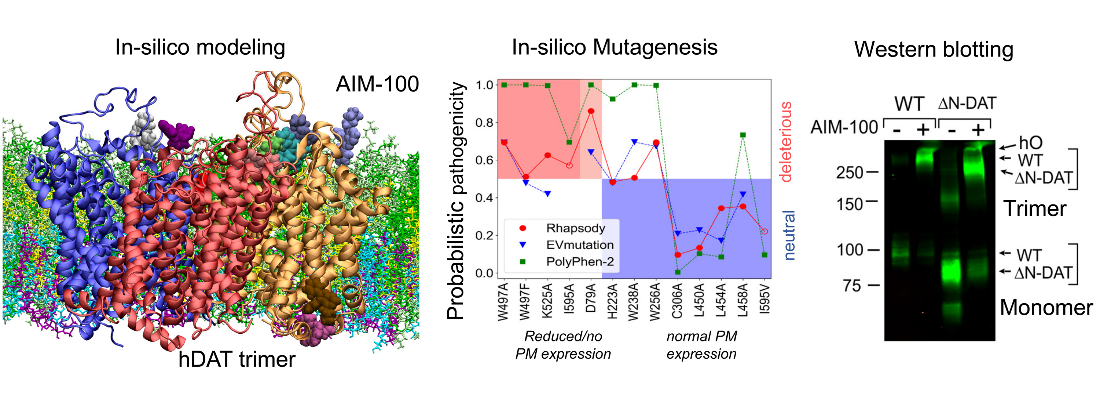
The Sorkin lab recently found that a furopyrimidine, AIM-100 triggered oligomerization of dopamine transporters (DATs), which promotes endocytosis and thereby may moderate dopaminergic transmission. Despite the significance of these events in mediating cellular responses, the underlying molecular mechanisms remain unclear. In the present study, we determined in-silico three structural models, for possible trimerization of DATs, in accord with the versatility of LeuT fold to stabilize dimeric or higher order constructs with a variety of packing geometries. Site-directed mutagenesis was performed to examine the effect of these elements/sites on DAT oligomerization and endocytosis (enhanced by AIM-100), and the experimental data were further interpreted using a novel machine learning classifier Rhapsody for assessing the impact of single amino acid variants (SAVs). The study suggests the possibility of controlling dopamine transport upon altering the oligomerization state of DAT by small molecular binding, as a possible intervention strategy to modulate dopaminergic signaling.
Cheng MH, Ponzoni L., Sorkina T., Lee JY, Zhang S., Sorkin A. & Bahar I. (2019) Trimerization of dopamine transporter triggered by AIM-100 binding: Underlying molecular mechanisms and effect of mutations, Neuropharmacology
Pre-post synaptic alignment through neuroligin-1 tunes synaptic transmission efficiency (Haas, Compans, Letellier, Bartol, et al. eLIFE. 2019)
 The nanoscale organization of neurotransmitter receptors regarding pre-synaptic release sites is a fundamental determinant of the synaptic transmission amplitude and reliability. How modifications in the pre- and post-synaptic machinery alignments affects synaptic currents, has only been addressed with computer modelling. Using single molecule super-resolution microscopy, we found a strong spatial correlation between AMPA receptor (AMPAR) nanodomains and the post-synaptic adhesion protein neuroligin-1 (NLG1). Expression of a truncated form of NLG1 disrupted this correlation without affecting the intrinsic AMPAR organization, shifting the pre-synaptic release machinery away from AMPAR nanodomains. Electrophysiology in dissociated and organotypic hippocampal rodent cultures shows these treatments significantly decrease AMPAR-mediated miniature and EPSC amplitudes. Computer modelling predicts that ~100 nm lateral shift between AMPAR nanoclusters and glutamate release sites induces a significant reduction in AMPAR-mediated currents. Thus, our results suggest the synapses necessity to release glutamate precisely in front of AMPAR nanodomains, to maintain a high synaptic responses efficiency.
The nanoscale organization of neurotransmitter receptors regarding pre-synaptic release sites is a fundamental determinant of the synaptic transmission amplitude and reliability. How modifications in the pre- and post-synaptic machinery alignments affects synaptic currents, has only been addressed with computer modelling. Using single molecule super-resolution microscopy, we found a strong spatial correlation between AMPA receptor (AMPAR) nanodomains and the post-synaptic adhesion protein neuroligin-1 (NLG1). Expression of a truncated form of NLG1 disrupted this correlation without affecting the intrinsic AMPAR organization, shifting the pre-synaptic release machinery away from AMPAR nanodomains. Electrophysiology in dissociated and organotypic hippocampal rodent cultures shows these treatments significantly decrease AMPAR-mediated miniature and EPSC amplitudes. Computer modelling predicts that ~100 nm lateral shift between AMPAR nanoclusters and glutamate release sites induces a significant reduction in AMPAR-mediated currents. Thus, our results suggest the synapses necessity to release glutamate precisely in front of AMPAR nanodomains, to maintain a high synaptic responses efficiency.
Haas KT, Compans B, Letellier M, Bartol TM, et al. (2019) Pre-post synaptic alignment through neuroligin-1 tunes synaptic transmission efficiency, eLIFE(2019)
Inferring the Assembly Network of Influenza Virus (Majarian, Murphy, Lakdawala, PLOS. 2019)
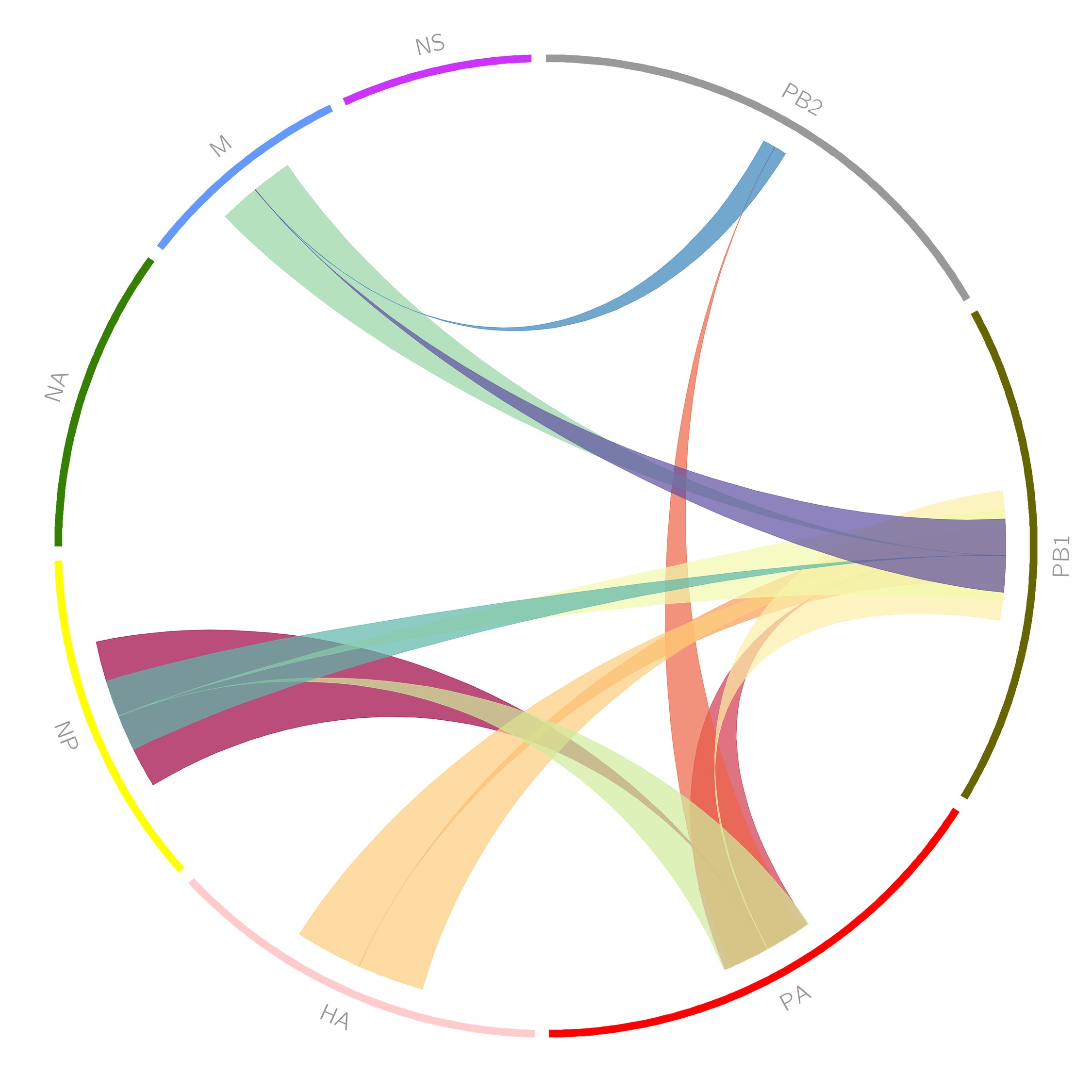 In influenza virus infected cells, eight viral genomic RNPs are assembled into progeny virions, which predominantly contain one copy of each. An important question is the order of the assembly of all eight RNPs. In an article in PLoS Computational Biology, MMBioS TR&D4 members Xiongto Ruan and Bob Murphy collaborated with Seema Lakdawala from the University of Pittsburgh School of Medicine to address this question. Using sets of fluorescence microscope images of four vRNPs, they estimated the likelihood that the subcellular distribution of one vRNP can be predicted from the distribution of one, two or three others. The figure shows the significant pairwise relationships that were learned (it was featured on the cover of the January 2019 issue of the journal). These pairwise and higher order relationships were used to infer the assembly network for all eight vRNPs.
In influenza virus infected cells, eight viral genomic RNPs are assembled into progeny virions, which predominantly contain one copy of each. An important question is the order of the assembly of all eight RNPs. In an article in PLoS Computational Biology, MMBioS TR&D4 members Xiongto Ruan and Bob Murphy collaborated with Seema Lakdawala from the University of Pittsburgh School of Medicine to address this question. Using sets of fluorescence microscope images of four vRNPs, they estimated the likelihood that the subcellular distribution of one vRNP can be predicted from the distribution of one, two or three others. The figure shows the significant pairwise relationships that were learned (it was featured on the cover of the January 2019 issue of the journal). These pairwise and higher order relationships were used to infer the assembly network for all eight vRNPs.
Majarian T, Murphy R, Lakdawala S (2019) Evaluation of methods for generative modeling of cell and nuclear shape, PLOS (2019)
2018 Highlights
PINK1 Interacts with VCP/p97 and Activates PKA to Promote NSFL1C/p47 Phosphorylation and Dendritic Arborization in Neurons (eNeuro. 2018)
 We found that PINK1 binds and phosphorylates the catalytic subunit of PKA at T197 [PKAcat(pT197)], a site known to activate the PKA holoenzyme. PKA in turn phosphorylates p47 at a novel site (S176) to regulate dendritic complexity. Given that PINK1 physically interacts with both the PKA holoenzyme and the VCP-p47 complex to promote dendritic arborization, we propose that PINK1 scaffolds a novel PINK1-VCP-PKA-p47 signaling pathway to orchestrate dendritogenesis in neurons. These findings highlight an important mechanism by which proteins genetically implicated in Parkinson’s disease (PD; PINK1) and frontotemporal dementia (FTD; VCP) interact to support the health and maintenance of neuronal arbors.
We found that PINK1 binds and phosphorylates the catalytic subunit of PKA at T197 [PKAcat(pT197)], a site known to activate the PKA holoenzyme. PKA in turn phosphorylates p47 at a novel site (S176) to regulate dendritic complexity. Given that PINK1 physically interacts with both the PKA holoenzyme and the VCP-p47 complex to promote dendritic arborization, we propose that PINK1 scaffolds a novel PINK1-VCP-PKA-p47 signaling pathway to orchestrate dendritogenesis in neurons. These findings highlight an important mechanism by which proteins genetically implicated in Parkinson’s disease (PD; PINK1) and frontotemporal dementia (FTD; VCP) interact to support the health and maintenance of neuronal arbors.
Wang KZQ, Steer E, Otero PA, Bateman NW, Cheng MH, Scott AL, Wu C, Bahar I, Shih YT, Hsueh YP and Chu CT (2018) PINK1 Interacts with VCP/p97 and Activates PKA to Promote NSFL1C/p47 Phosphorylation and Dendritic Arborization in Neurons, eNeuro (2018)
Improved methods for modeling cell shape (Ruan, Murphy, Bioinformatics. 2018)
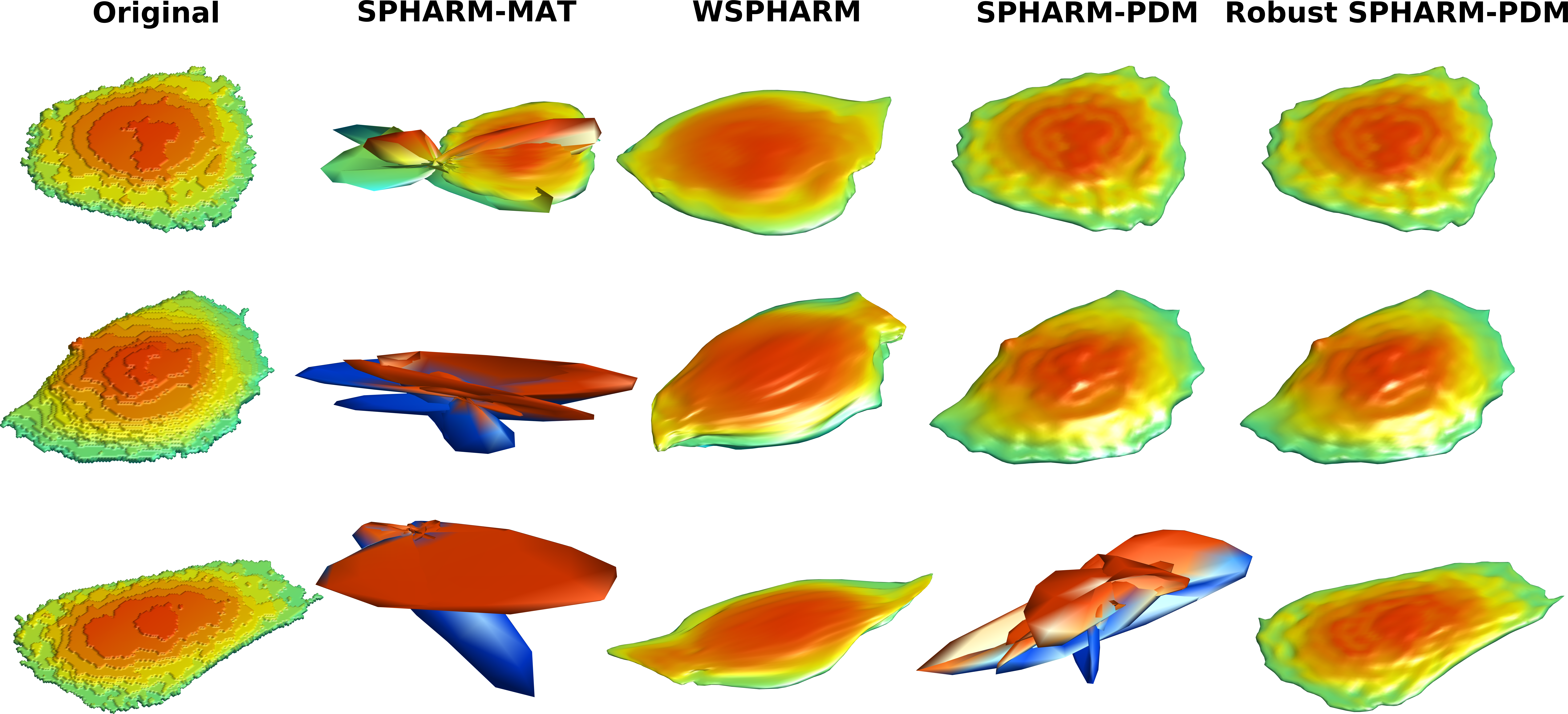 In a recent paper in Bioinformatics, Xiongtao Ruan and Bob Murphy of TR&D4 addressed the question of how best to model cell and nuclear shape. They compared a number of previously described methods for constructing models from either 2D or 3D images, including both traditional and deep learning methods. The comparison was done with respect to how accurately a lower dimensional shape representation in a model could be used to reconstruct the shapes with which it was trained. This unsupervised task is much more demanding than the more common supervised task of distinguishing between different classes of cells. The results showed that outline PCA performed best for 2D images but that none of the previous methods worked well for 3D shapes. The best of those previous methods were based on spherical harmonics, but common implementations often failed at producing a good representation of a cell (see figure). They identified the causes and introduced a robust method that worked well in all case, including for highly non-spherical cells like neurons. The code for these methods is available in the latest release of CellOrganizer.
In a recent paper in Bioinformatics, Xiongtao Ruan and Bob Murphy of TR&D4 addressed the question of how best to model cell and nuclear shape. They compared a number of previously described methods for constructing models from either 2D or 3D images, including both traditional and deep learning methods. The comparison was done with respect to how accurately a lower dimensional shape representation in a model could be used to reconstruct the shapes with which it was trained. This unsupervised task is much more demanding than the more common supervised task of distinguishing between different classes of cells. The results showed that outline PCA performed best for 2D images but that none of the previous methods worked well for 3D shapes. The best of those previous methods were based on spherical harmonics, but common implementations often failed at producing a good representation of a cell (see figure). They identified the causes and introduced a robust method that worked well in all case, including for highly non-spherical cells like neurons. The code for these methods is available in the latest release of CellOrganizer.
Ruan X, Murphy R (2018) Learning the sequence of influenza A genome assembly during viral replication using point process models and fluorescence in situ hybridization, Bioinformatics (2018)
Structural dynamics is a determinant of the functional significance of missense variants (Ponzoni, Bahar, PNAS. 2018)
 We demonstrated that the analysis of a protein’s intrinsic dynamics can be successfully used to improve the prediction of the effect of point mutations on a protein functionality. This method employs ANM/GNM tools implemented in the ProDy Python package, in conjunction with other well-established predictors based on amino acid conservation/coevolution and structural properties of the mutation site (e.g. solvent-accessible surface area).
We demonstrated that the analysis of a protein’s intrinsic dynamics can be successfully used to improve the prediction of the effect of point mutations on a protein functionality. This method employs ANM/GNM tools implemented in the ProDy Python package, in conjunction with other well-established predictors based on amino acid conservation/coevolution and structural properties of the mutation site (e.g. solvent-accessible surface area).
Ponzoni L, Bahar I (2018) Structural dynamics is a determinant of the functional significance of missense variants, PNAS (2018)
2017 Highlights
Development of a new methodology for investigating the structural dynamics of chromatin and its relation to gene expression and regulation (Sauerwald, Zheng, Kingsford & Bahar, Nucleic Acids Res. 2017), driven by DBP6
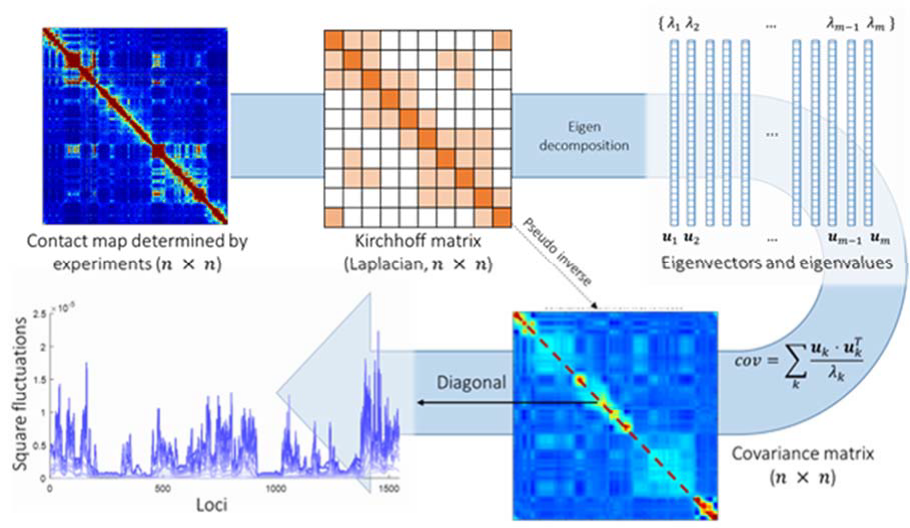 3D genome organization, but we still lack an understanding of the structural dynamics of chromosomes. We adapted our well-established protein-modeling framework, the Gaussian Network Model (GNM), to model chromatin dynamics using Hi-C data. We used GNM to identify spatial couplings at multiple scales: it can quantify the correlated fluctuations in the Advances in Hi-C technology now permit us to study positions of gene loci, find large genomic compartments and smaller topologically-associating domains (TADs) that undergo en-bloc movements, and identify dynamically coupled distal regions along the chromosomes (Fig 1). We found that the predictions of the GNM correlate well with genome- wide experimental measurements. We used the GNM to identify novel cross-correlated distal domains (CCDDs) representing pairs of regions distinguished by their long-range dynamic coupling and found that CCDDs are associated with increased gene co-expression.
3D genome organization, but we still lack an understanding of the structural dynamics of chromosomes. We adapted our well-established protein-modeling framework, the Gaussian Network Model (GNM), to model chromatin dynamics using Hi-C data. We used GNM to identify spatial couplings at multiple scales: it can quantify the correlated fluctuations in the Advances in Hi-C technology now permit us to study positions of gene loci, find large genomic compartments and smaller topologically-associating domains (TADs) that undergo en-bloc movements, and identify dynamically coupled distal regions along the chromosomes (Fig 1). We found that the predictions of the GNM correlate well with genome- wide experimental measurements. We used the GNM to identify novel cross-correlated distal domains (CCDDs) representing pairs of regions distinguished by their long-range dynamic coupling and found that CCDDs are associated with increased gene co-expression.
Elucidation of anion conductance mechanism of human excitatory amino acid transporters and coupling to glutamate transport (driven by DBP1; published in Cheng, Torres-Salazar, Gonzalez-Suarez, Amara, and Bahar, ELife 2017).
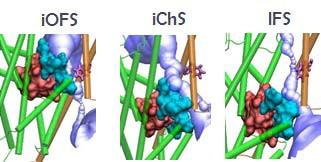 We identified an intermediate anion channeling state (iChS) during the global transition from the outward facing (OF) to inward facing state (IFS). Our prediction was tested and validated by experimental study conducted in the Amara lab (NIMH). Critical residues and interactions were analyzed by SCAM, electrophysiology and substrate uptake experiments
We identified an intermediate anion channeling state (iChS) during the global transition from the outward facing (OF) to inward facing state (IFS). Our prediction was tested and validated by experimental study conducted in the Amara lab (NIMH). Critical residues and interactions were analyzed by SCAM, electrophysiology and substrate uptake experiments
Development of an integrative framework for combining structural and kinetic data and computing technology at multiple scales.
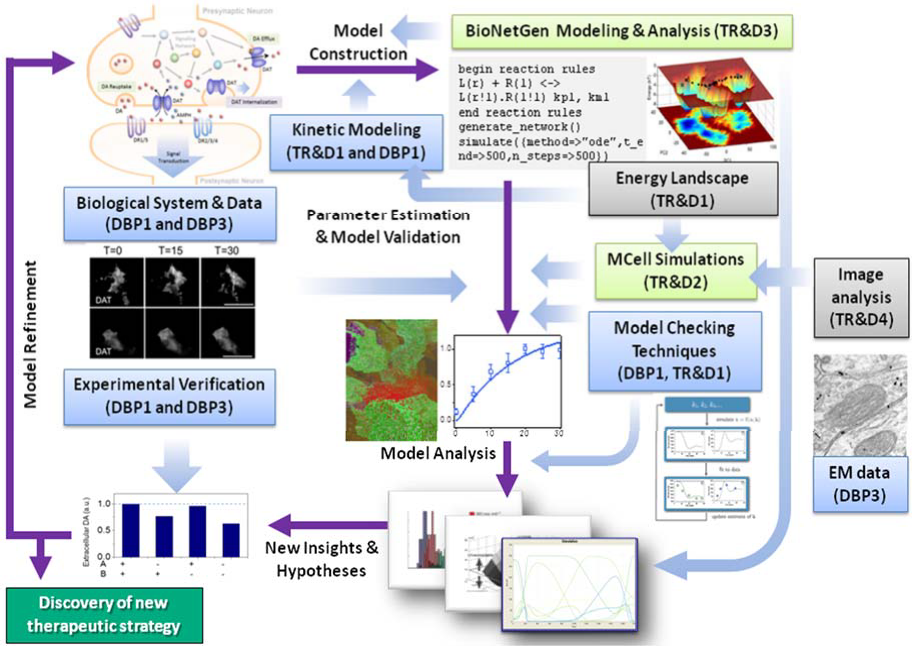 We have made progress in integrating the different software tools that we are developing in a suitable pipeline. One first step was to facilitate and further accelerate this goal by the redefinition of existing DBPs (DBP1-4) and design of the new DBPs (DBP6-8) to contain each technological challenges that require the involvement of at least two of the TR&Ds. TR&D team members driven by the same DBP would naturally join forces and integrate their tools to respond to the needs of the DBP. An example of ongoing integrated project is illustrated in Figure 3. This project, focused on simulating dopaminergic transmission driven by DBP1 and DBP2 combines ProDy, MCell, and BioNetGen developed by the respective TR&D1, 2 and 3 of the new term; and it will be further developed to automate image analyses with the involvement of Bioimage Analysis TR&D (TR&D3 in Year 5, TR&D4 in new term). A first paper on this topic, co-authored by several MMBioS TR&D leaders has been submitted for publication in eNeuro, and a revised version has been recently submitted. TR&D numbers in the caption below refer to the new term TR&Ds.
We have made progress in integrating the different software tools that we are developing in a suitable pipeline. One first step was to facilitate and further accelerate this goal by the redefinition of existing DBPs (DBP1-4) and design of the new DBPs (DBP6-8) to contain each technological challenges that require the involvement of at least two of the TR&Ds. TR&D team members driven by the same DBP would naturally join forces and integrate their tools to respond to the needs of the DBP. An example of ongoing integrated project is illustrated in Figure 3. This project, focused on simulating dopaminergic transmission driven by DBP1 and DBP2 combines ProDy, MCell, and BioNetGen developed by the respective TR&D1, 2 and 3 of the new term; and it will be further developed to automate image analyses with the involvement of Bioimage Analysis TR&D (TR&D3 in Year 5, TR&D4 in new term). A first paper on this topic, co-authored by several MMBioS TR&D leaders has been submitted for publication in eNeuro, and a revised version has been recently submitted. TR&D numbers in the caption below refer to the new term TR&Ds.
Large scale visualization of rule-based models.
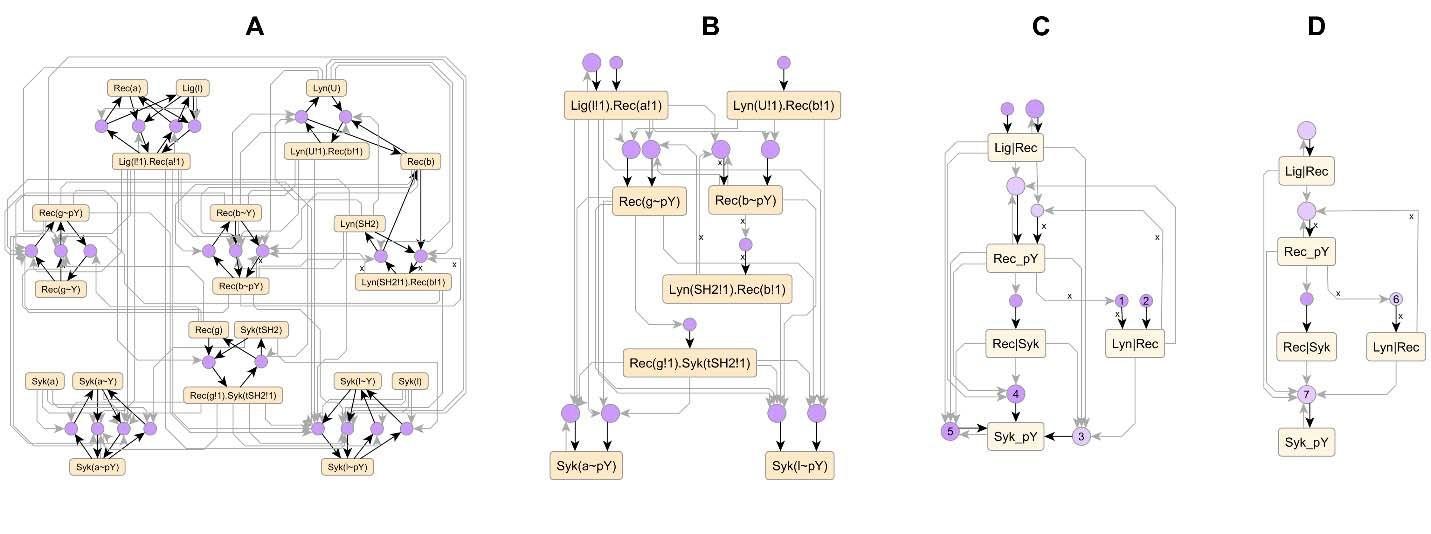 Signaling in living cells is mediated through a complex network of chemical interactions. Current predictive models of signal pathways have hundreds of reaction rules that specify chemical interactions, and a comprehensive model of a stem cell or cancer cell would be expected to have many more. Visualizations of rules and their interactions are needed to navigate, organize, communicate and analyze large signaling models. In this work, we have developed: (i) a novel visualization for individual rules that compactly conveys what each rule does, (ii) a comprehensive visualization of a set of rules as a network of regulatory interactions called an atom-rule (AR) graph, and (iii) a set of procedures for compressing the AR graph into a pathway diagram that highlights underlying signaling motifs such as feedback and feed-forward loops. We show that these visualizations are compact and informative across models of widely varying sizes. The methods developed here not only improve the understandability of current models, but also establish principles for organizing the much larger models of the future.
Signaling in living cells is mediated through a complex network of chemical interactions. Current predictive models of signal pathways have hundreds of reaction rules that specify chemical interactions, and a comprehensive model of a stem cell or cancer cell would be expected to have many more. Visualizations of rules and their interactions are needed to navigate, organize, communicate and analyze large signaling models. In this work, we have developed: (i) a novel visualization for individual rules that compactly conveys what each rule does, (ii) a comprehensive visualization of a set of rules as a network of regulatory interactions called an atom-rule (AR) graph, and (iii) a set of procedures for compressing the AR graph into a pathway diagram that highlights underlying signaling motifs such as feedback and feed-forward loops. We show that these visualizations are compact and informative across models of widely varying sizes. The methods developed here not only improve the understandability of current models, but also establish principles for organizing the much larger models of the future.
Integration of MCellR into MCell/CellBlender validated using a model of SynGAP/PSD95 interactions in the PSD driven by DPB2.
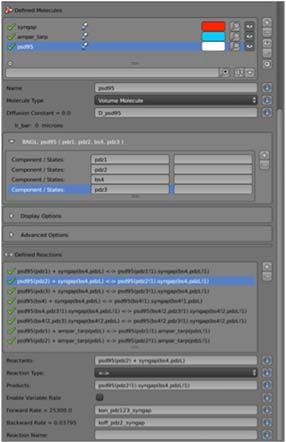 The spatial biochemical models of SynGAP/PSD95 interactions in the PSD studied by the Kennedy Lab (DBP2) require the new spatial rule-based modeling methods being developed in TR&D2. A major accomplishment of the past year is merging the MCellR code-base with the main MCell code-base, making this sophisticated technology easily accessible to the user through the MCell/CellBlender GUI (Fig 5 left panel), and validating the utility and correctness of MCellR through implementation of a model of synGAP/PSD95 cross-linking interactions (Fig 5 right panel). We also observed a 4X speedup in model execution speed over our previous simple implementation of complex.
The spatial biochemical models of SynGAP/PSD95 interactions in the PSD studied by the Kennedy Lab (DBP2) require the new spatial rule-based modeling methods being developed in TR&D2. A major accomplishment of the past year is merging the MCellR code-base with the main MCell code-base, making this sophisticated technology easily accessible to the user through the MCell/CellBlender GUI (Fig 5 left panel), and validating the utility and correctness of MCellR through implementation of a model of synGAP/PSD95 cross-linking interactions (Fig 5 right panel). We also observed a 4X speedup in model execution speed over our previous simple implementation of complex.
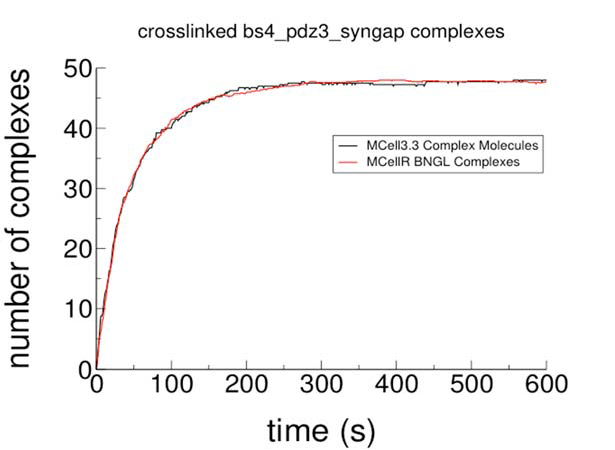
Development of approaches for identifying potential causal relationships between the spatial distributions of different proteins during T cell signaling.
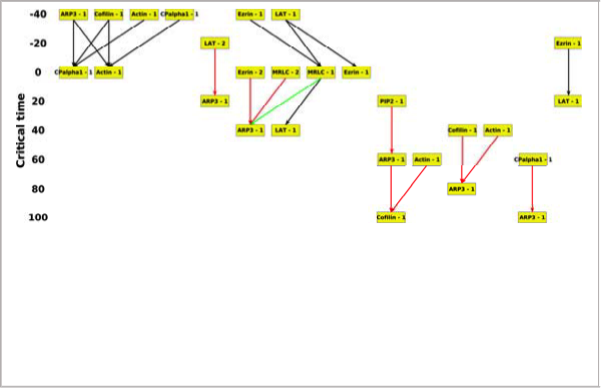 The idea is to identify a relationship in which a change in the
The idea is to identify a relationship in which a change in the
concentration of one protein in one cell region consistently is associated with a change in the concentration of another protein in the same or a different region. We used the data from our Science Signaling paper reported last year to construct a model for T cells undergoing stimulation by both the T cell receptor and the costimulatory receptor. The model recapitulated known relationships and identified new one. Most important, the model learned from these costimulatory conditions was predictive of the protein dynamics in different cells under conditions in which the costimulatory receptor was blocked (data not used in training the model). The paper describing this work was accepted for presentation at ISMB 2017 (one of the top two computational biology conferences, with an acceptance rate of only 16%) and published in Bioinformatics.
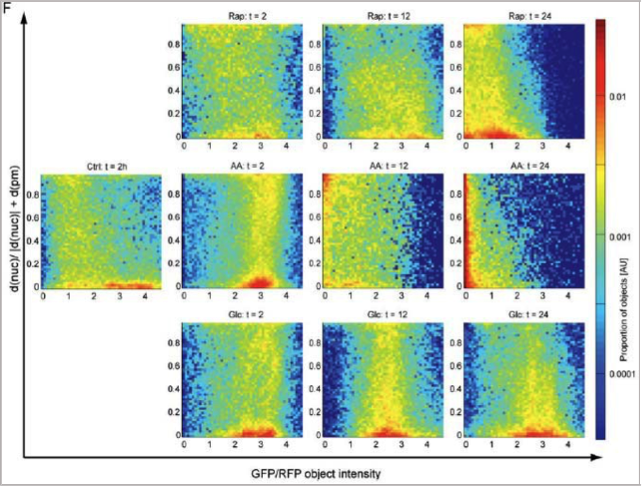
The second figure (right) is from the collaboration with Jörn Dengjel on C&SP29. We used our image analysis methods to measure the acidification and transport of autophagosomes under different cell starvation conditions. The results showed that processing was different under three conditions: in one case, acidification and transport were blocked, in another, neither were blocked, and in the last, transport but not acidification was blocked. The paper was published in Autophagy with Figure YY as the cover image.
2016 Highlights
Modeling with BioNetGen Gives MMBioS Team Insight into How Immune System Decides to Attack — or Let Be
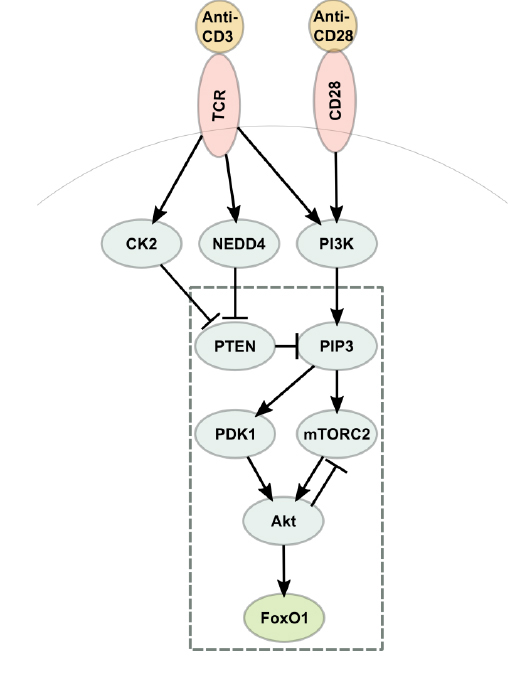 Friend or Foe?
Friend or Foe?
A mix of computer modeling and laboratory experiments has helped reveal how the body differentiates “friend from foe.” Using their BioNetGen computer tool for simulating biochemistry, MMBioS members and colleagues have painted a sharper picture of how T cells, the advance scouts of the immune system, decide when to protect bodily tissues from immune attack—and when to lead the attack. The finding may guide future efforts to control human diseases like diabetes and cancer.
When T cells—a type of white blood cell—encounter other cells in the body, “they have to decide what kind of a response to make,” says James Faeder, project co-leader of MMBioS’s Technology Research and Development Project 2 Team and associate professor of computational & systems biology, University of Pittsburgh. “Is that a threat, or is it something benign? Depending on their assessment of a possible threat they can either become activated, immune-boosting cells that kill pathogens, or they can tamp down those responses.”
See more about MMBioS research in cell modeling.
Tools for determining the spatial relationships between different cell components
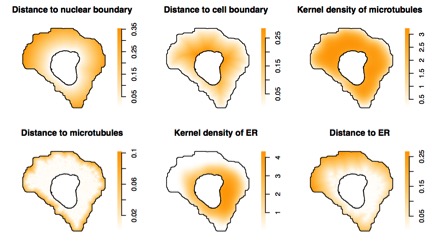
An important task for understanding how cells are organized is determining which components have spatial patterns that are related to each other. We can think of this as learning the “order of assembly” of cells – which parts affect the patterns of other parts. We added tools for analyzing this to CellOrganizer and used them to demonstrate that different organelles that show “punctate” patterns (such as peroxisomes and coated pits) can be distinguished from each other by measuring how much their spatial patterns depend upon microtubules1. We further extended this to perform rigorous statistical testing of the extent to which these organelles depend upon four different components: the cell membrane, nuclear membrane, endoplasmic reticulum membrane, and microtubules2.
[Image Left] The panels show maps of different factors that might influence punctate organelle distribution. The factors are calculated from images of cells labeled with probes for DNA, microtubules, and endoplasmic reticulum. The values of each factor at each position in a typical cell are shown.
1Johnson GR, Buck TE, Sullivan DP, Rohde GK, Murphy RF Joint modeling of cell and nuclear shape variationMolecular biology of the cell26(22): 4046-56. doi: 10.1091/mbc.E15-06-0370.
2Li Y, Majarian TD, Naik AW, Johnson GR, Murphy RF (2016) Point process models for localization and interdependence of punctate cellular structures Cytometry Part A89:in press. doi: 10.1002/cyto.a.22873.
See more about MMBioS research in image processing
Pipeline for creation of spatiotemporal maps of T cell signaling proteins
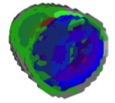

Using a combination of diffeomorphic methods and improved cell segmentation, we developed a CellOrganizer pipeline for use in DPB4 to construct models of the 4D distributions of actin and 8 of its regulators during the response of T cells to antigen presentation. This work, published in Science Signaling (Roybal et al) revealed that WAVE2 and cofilin play a critical role in co-stimulation through CD28, and this was confirmed by reconstitution experiments. Spatiotemporal maps of all nine proteins were made publically available (http://murphylab.web.cmu.edu/data/TcellModels/) along with movies of the concentration of three representative proteins in 3D over time as T cell costimulation proceeds either as 3D projections.
(Figure right shows the first frame) (View full movie) or (Central 2d slices).
Frame from movie showing 4D distributions of cofilin (red), MRLC (green) and WAVE2 (blue). At this timepoint, cofilin and WAVE2 have translocated to the synapse but MRLC is lagging.
Roybal KT, Buck TE, Ruan X, Cho BH, Clark DJ, Ambler R, et al. (2016) Computational spatiotemporal analysis identifies WAVE2 and cofilin as joint regulators of costimulation-mediated T cell actin dynamics. Science Signaling. 9(424):1-13. doi: 10.1126/scisignal.aad4149.
See more about MMBioS research in image processing
New Release of iGNM Database
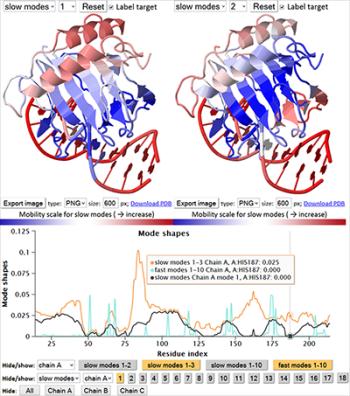
Gaussian network model (GNM) is a simple yet powerful model for investigating the intrinsic dynamics of proteins and their complexes. Intrinsic dynamics refers to conformational changes intrinsically favored by 3D structure, which often underlie the adaptation of biomolecules to functional interactions. Which structural elements undergo large fluctuations, or which ones provide adequate flexibility to enable conformational changes (e.g. hinge-bending sites) that may be relevant to function? Furthermore, which structural elements are subject to strongly correlated (or anticorrelated) motions, toward gaining insights into allosterically coupled regions? The GNM addresses these questions. It further allows to dissect these properties into the contributions of individual modes, thus elucidating the cooperative couplings underlied by low frequency modes.
 The updated iGNM 2.0 covers more than 95% of the structures in the Protein Data Bank (PDB). Advanced search and visualization capabilities, both 2D and 3D, permit users to retrieve information on inter-residue and inter-domain cross-correlations, cooperative modes of motion, the location of hinge sites and energy localization spots. A unique feature of iGNM 2.0 is the accessibility of data for biological assemblies (BAs), rather than the single chains or asymmetric units (Asym), thus providing insights into the dynamic properties of biologically functional entities.
The updated iGNM 2.0 covers more than 95% of the structures in the Protein Data Bank (PDB). Advanced search and visualization capabilities, both 2D and 3D, permit users to retrieve information on inter-residue and inter-domain cross-correlations, cooperative modes of motion, the location of hinge sites and energy localization spots. A unique feature of iGNM 2.0 is the accessibility of data for biological assemblies (BAs), rather than the single chains or asymmetric units (Asym), thus providing insights into the dynamic properties of biologically functional entities.
Examination of structures of even 104 residues in iGNM 2.0 showed that the accuracy of the results did not decrease with increasing size (N).
Li,H., Chang,YY, Yang,LW and Bahar,I. (2016) iGNM 2.0: The Gaussian Network Model Database for Biomolecular Structural Dynamics. Nucleic Acids Res., 44, D415-D422.
See more about MMBioS research in molecular modeling
Improved Sampling of Cell-Scale Models using the Weighted Ensemble Strategy
 The “weighted ensemble” (WE) strategy for orchestrating a large set of parallel simulations has been established as an effective tool for efficiently calculating kinetic and equilibrium observables in molecular systems – and now has been extended to spatially resolved cell-scale systems by MMBioS researchers. In a collaboration among several MMBioS groups, the WESTPA implementation of WE has been used to control MCell simulations, including models built using a BioNetGen-CellOrganizer pipeline for situating complex biochemistry within spatially realistic cell models. As sketched, the WE strategy enables computational effort to be focused on rare, difficult-to-sample events – essentially increasing the precision of measurements in the tails of distributions.
The “weighted ensemble” (WE) strategy for orchestrating a large set of parallel simulations has been established as an effective tool for efficiently calculating kinetic and equilibrium observables in molecular systems – and now has been extended to spatially resolved cell-scale systems by MMBioS researchers. In a collaboration among several MMBioS groups, the WESTPA implementation of WE has been used to control MCell simulations, including models built using a BioNetGen-CellOrganizer pipeline for situating complex biochemistry within spatially realistic cell models. As sketched, the WE strategy enables computational effort to be focused on rare, difficult-to-sample events – essentially increasing the precision of measurements in the tails of distributions.  In an application to an MCell model of the frog neuro-muscular junction (NMJ), WE simulation enabled calculation of vesicle release probabilities under particularly challenging low-calcium concentrations that could not be well sampled by conventional MCell simulation: see red-circled points in the figure. This in turn, enabled validation of the model by confirmation of an established NMJ empirical power-law relationship between calcium-concentration and release probability. WE yielded estimates of observables in less overall computing time than would be required in ordinary parallelization, thus exhibiting super-linear parallel performance.
In an application to an MCell model of the frog neuro-muscular junction (NMJ), WE simulation enabled calculation of vesicle release probabilities under particularly challenging low-calcium concentrations that could not be well sampled by conventional MCell simulation: see red-circled points in the figure. This in turn, enabled validation of the model by confirmation of an established NMJ empirical power-law relationship between calcium-concentration and release probability. WE yielded estimates of observables in less overall computing time than would be required in ordinary parallelization, thus exhibiting super-linear parallel performance.
Donovan RM, Tapia JJ, Sullivan DP, Faeder JR, Murphy RF, Dittrich M, Zuckerman DM (2016). Unbiased Rare Event Sampling in Spatial Stochastic Systems Biology Models Using a Weighted Ensemble of Trajectories PLoS Comput Biol. 12(2):e1004611
See more about MMBioS research in molecular modeling
Anatomy and Function of an Excitatory Network in the Visual Cortex

MMBioS researcher Greg Hood’s collaboration with Wei-Chung Allen Lee of Harvard University and R. Clay Reid of the Allen Institute for Brain Science concerning the reconstruction of an excitatory nerve-cell network in the mouse brain cortex at a subcellular level using the AlignTK software has been published in Nature.
Lee WA, Bonin V, Reed M, Graham BJ, Hood G, Glattfelder K, Reid RC (2016) Anatomy and function of an excitatory network in the visual cortex Nature 532: 370–374.
See more about MMBioS research in image processing
Memory Capacity of Brain 10 Times More than Previously Thought
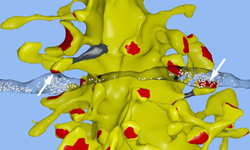
Salk researchers and MMBioS team members Tom Bartol and Terry Sejnowski, along with their collaborators, have achieved critical insight into the size of neural connections, putting the memory capacity of the brain far higher than common estimates. The new work also answers a longstanding question as to how the brain is so energy efficient and could help engineers build computers that are incredibly powerful but also conserve energy.
In a computational reconstruction of brain tissue in the hippocampus, Salk scientists and UT-Austin scientists found the unusual occurrence of two synapses from the axon of one neuron (translucent black strip) forming onto two spines on the same dendrite of a second neuron (yellow). Separate terminals from one neuron’s axon are shown in synaptic contact with two spines (arrows) on the same dendrite of a second neuron in the hippocampus. The spine head volumes, synaptic contact areas (red), neck diameters (gray) and number of presynaptic vesicles (white spheres) of these two synapses are almost identical.
Credit: Salk Institute
TM Bartol, C Bromer, J Kinney, MA Chirillo, JN Bourne, KM Harris, TJ Sejnowski (2015) Nanoconnectomic Upper Bound on the Variability of Synaptic Plasticity. eLife 2015;4:e10778.
See more about MMBioS research in cell modeling.
Development and improvements to MCell
The MCell modeling and simulation platform provides the core capabilities for spatial simulation reaction-diffusion dynamics at complex biological interfaces. Highlights of progress include:
libMCell API We have developed a high-level set of routines, which enable the creation and simulation of MCell models via API calls alone, i.e., without invoking any parsed MDL description of the model. Current functionality includes creation of species, defining reactions, defining molecule releases, creating model geometry, and simulation and runtime control.
MCell testing framework This framework, called nutmeg (https://github.com/haskelladdict/nutmeg), has completely replaced our previous, python based unit test framework. During the year we made nutmeg much more robust and added many additional test cases.
Implementation of new simulation capabilities that extend the range of simulations that can be performed in order to address important biological questions, such as the how membrane dynamics and signal transduction interact or how multisite phosphorylation, binding, and cooperativity affect signaling. Highlights include:
- Dynamic geometries Users can now create dynamical meshes using Blender and use these to drive an MCell simulation of reaction and diffusion with moving boundaries.
- Modeling pipeline Users can build complex biochemical models using BioNetGen and sample complex cellular geometries from imaging data using CellOrganizer and combine those into a single model that can be simulated in CellBlender (in collaboration with TRD3).
- Recovery of protein-protein interactions and other aspect of biochemical mechanisms from reaction network models using Atomizer. The automated conversion of these models into a rule-based format allows for comparative analysis of models and reuse of existing model components.
- Visualization of regulatory structurein rule-based models New visualization methods have been developed that enable global visualization of models exhibiting combinatorial complexity.
D. P. Sullivan, J. J. Tapia, R. Arepally, J. Czech, R. F. Murphy, M. Dittrich, and J. R. Faeder, “Design Automation for Biological Models : A Pipeline that Incorporates Spatial and Molecular Complexity,” in 25th edition of the Great Lakes Symposium on VLSI, 2015, pp. 321–323.
J. A. P. Sekar, J.-J. Tapia, and J. R. Faeder, “Visualizing Regulation in Rule-based Models.” Submitted to Bioinformatics . arXiv:1509.00896 [q-bio.QM].
See more about MMBioS research in cell modeling.
Development of CellBlender
CellBlender is a graphical interface for model construction, simulation, and analysis of complex spatial models of reaction-diffusion systems Highlights of progress include
- A major redesign of the interface to consolidate functionality based on a single toolbar.
- Implementation of a robust parameter handling system
- Functionality for running simulations from within the interface taking advantage of parallel computing resources allowing users to manage on-going simulation streams.
- Direct access to run-time error logs within the Blender interface.
- Model and geometry importers that allow import of compartmental reaction network models in either SBML or compartmental BNGL format.
- Implementation of internal data model provides backward compatibility with future CellBlender versions.
- Release of CellBlender version 1.0 and publication of an article featuring MCell and CellBlender in the Encyclopedia of Computational Neuroscience
T. Bartol, M. Dittrich, and J. Faeder, “MCell,” in Encyclopedia of Computational Neuroscience, D. Jaeger and R. Jung, Eds. Springer New York, 2014, pp. 1–5.
See more about MMBioS research in cell modeling.
2015 Highlights
A Mutation in Transmembrane Domain 7 (TM7) of Excitatory Amino Acid Transporters Disrupts the Substrate-dependent Gating of the Intrinsic Anion Conductance and Drives the Channel into a Constitutively Open State
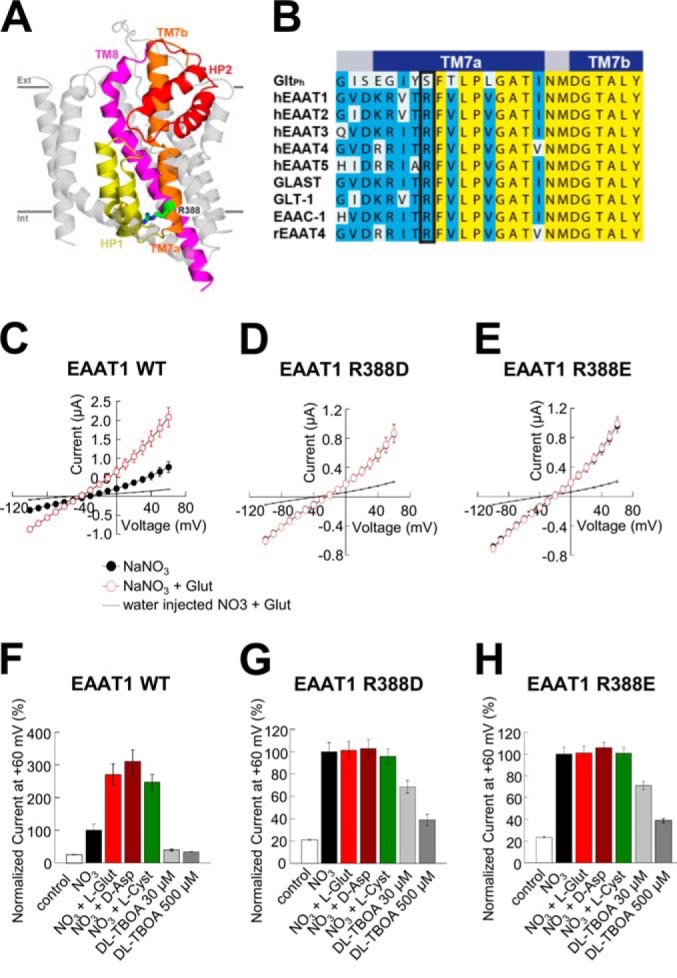
In the mammalian central nervous system, excitatory amino acid transporters (EAATs) are responsible for the clearance of glutamate after synaptic release. This energetically demanding activity is crucial for precise neuronal communication and for maintaining extracellular glutamate concentrations below neurotoxic levels. In addition to their ability to recapture glutamate from the extracellular space, EAATs exhibit a sodium and glutamate gated anion conductance. Here we show that substitution of a conserved positively charged residue (Arg388, hEAAT1) in transmembrane domain 7 with a negatively charged amino acid eliminates the ability of glutamate to further activate the anion conductance.
When expressed in oocytes, R388D or R388E mutants show large anion currents that display no further increase in amplitude after application of saturating concentrations of Na(+) and glutamate.
They also show a substantially reduced transport activity. The mutant transporters appear to exist preferentially in a sodium and glutamate independent constitutive open channel state that rarely transitions to complete the transport cycle. In addition, the accessibility of cytoplasmic residues to membrane permeant modifying reagents supports the idea that this substrate independent open state correlates with an intermediate outward facing conformation of the transporter. Our data provide additional insights into the mechanism by which substrates gate the anion conductance in EAATs and suggest that in EAAT1, Arg388 is a critical element for the structural coupling between the substrate translocation and the gating mechanisms of the EAATassociated anion channel.
TorresSalazar, D.; Jiang, J.; Divito, C. B.; GarciaOlivares, J.; Amara, S. G. A Mutation in Transmembrane Domain 7 (TM7) of Excitatory Amino Acid Transporters Disrupts the Substrate dependent Gating of the Intrinsic Anion Conductance and Drives the Channel into a Constitutiv ely Open State. J Biol Chem 2015, 290 (38), 2297722990.
Development of diffeomorphic and dynamic models of cell and nuclear shape
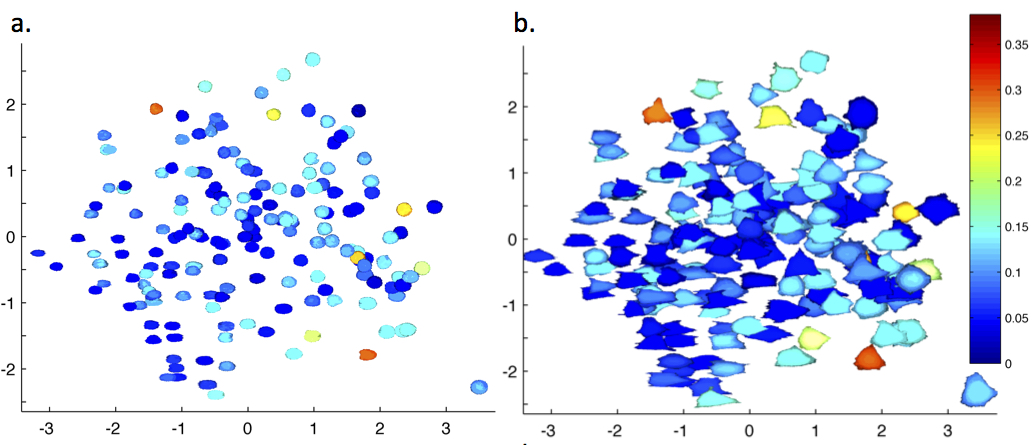
We added a major new capability to our open source CellOrganizer system, the ability to construct diffeomorphic models of cell and nuclear shape. The models were developed because of the needs of DBP4; to carry out initial benchmarking we applied them to publicly available datasets and published an extensive study (Johnson et al, Mol. Biol. Cell). We measured the statistical dependency of nuclear shape on cell shape (below), and showed that drug treatment and gene inhibition can alter that dependency.
Projections of cells (right) and nuclei (left) onto the 2D subspace of highest-variance components in 7-dim diffeomorphic shape space. Cell shapes (left) are colored by the p-value that the nuclear shape predicted from the cell shape is closer to the actual shape compared to that expected by randomly choosing a nuclear shape from all examples; and the nuclear shapes on the right are colored similarly for predicting cell shape. Most cell shapes are predicted well from nuclear shape and vice versa.
, , , , . (2015) Joint modeling of cell and nuclear shape variation. Molecular biology of the cell. 26(22):4046-56. doi: 10.1091/mbc.E15-06-0370.
Amphetamine activates Rho GTPase signaling to mediate dopamine transporter internalization and acute behavioral effects of amphetamine
Acute amphetamine (AMPH) exposure elevates extracellular dopamine through a variety of mechanisms that include inhibition of dopamine reuptake, depletion of vesicular stores, and facilitation of dopamine efflux across the plasma membrane. Recent work has shown that the DAT substrate AMPH, unlike cocaine and other nontransported blockers, can also stimulate endocytosis of the plasma membrane dopamine transporter (DAT). Here, we show that when AMPH enters the cytoplasm it rapidly stimulates DAT internalization through a dynamin-dependent, clathrin-independent process. This effect, which can be observed in transfected cells, cultured dopamine neurons, and midbrain slices, is mediated by activation of the small GTPase RhoA. Inhibition of RhoA activity with C3 exotoxin or a dominant-negative RhoA blocks AMPH-induced DAT internalization. These actions depend on AMPH entry into the cell and are blocked by the DAT inhibitor cocaine. AMPH also stimulates cAMP accumulation and PKA-dependent inactivation of RhoA, thus providing a mechanism whereby PKA- and RhoA-dependent signaling pathways can interact to regulate the timing and robustness of AMPH’s effects on DAT internalization. Consistent with this model, the activation of D1/D5 receptors that couple to PKA in dopamine neurons antagonizes RhoA activation, DAT internalization, and hyperlocomotion observed in mice after AMPH treatment. These observations support the existence of an unanticipated intracellular target that mediates the effects of AMPH on RhoA and cAMP signaling and suggest new pathways to target to disrupt AMPH action.
Wheeler, D. S.; Underhill, S. M.; Stolz, D. B.; Murdoch, G. H.; Thiels, E.; Romero, G.; Amara, S. G. Amphetamine activates Rho GTPase signaling to mediate dopamine transporter internalization and acute behavioral effects of amphetamine. Proc. Natl. Acad. Sci. U. S. A112 (51), E7138-E7147.
Energy landscape for LeuT
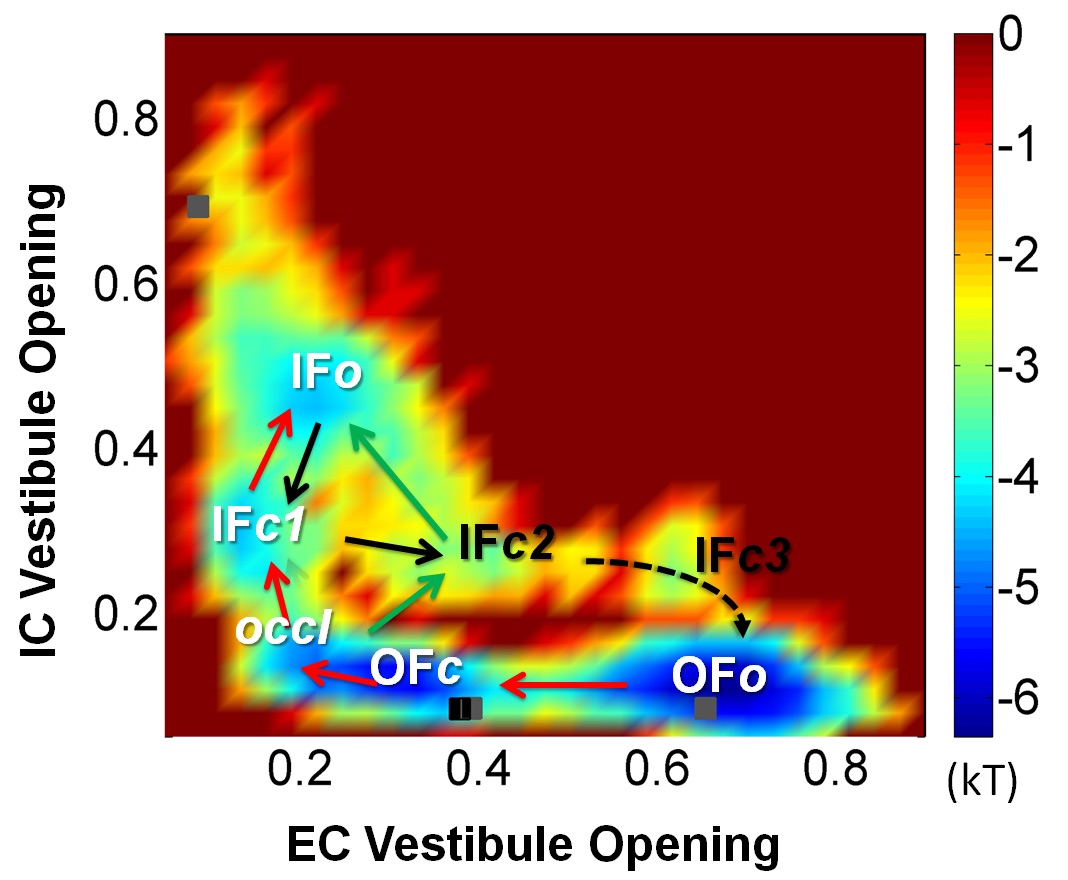
Development of a multiscale hybrid methodology for constructing LeuT energy landscape and delineating the molecular mechanism of substrate/neurotranmitter transport cycle by NSS family members The hybrid methodology, coMD, that we have recently developed [1] has been recently extended to construct the energy landscape near the functional states of LeuT (Fig 1) [2]. This is the first energy landscape constructed for this NSS family member. It is obtained by combining 10s of microseconds of Anton trajectories [3] and coMD sampling of transition regions. It permits to visualize the accessible states and transition paths.
Fig 1. Energy landscape for LeuT, sampled during its substrate transport cycle. OF and IF refer to outward-facing and inward-facing states, and each has multiple substates (o, c, etc.).
- References: 1. Gur M, Madura J, Bahar I. (2013) Biophys J 105: 164352. 2. Zomot E, Gur M, Bahar I. (2015) J Biol Chem 290: 54455. 3. Gur M, Zomot E, Cheng M, Bahar I. (2015) J Chem Phys 143: 243134
Insights into the Cooperative Dynamics of AMPAR
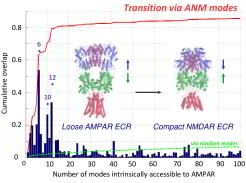 Interconversion from AMPAR to NMDAR conformation by moving in a space of soft modes. The dark blue bars show the overlap (correlation cosine) between ANM modes of AMPAR and the structural difference vector between AMPAR and NMDAR; the red curve displays the cumulative overlap; the dotted green curve shows the expected cumulative overlap if the modes were equally contributing each. The inset shows the initial structures of AMPAR (left) and NMDAR (right).
Interconversion from AMPAR to NMDAR conformation by moving in a space of soft modes. The dark blue bars show the overlap (correlation cosine) between ANM modes of AMPAR and the structural difference vector between AMPAR and NMDAR; the red curve displays the cumulative overlap; the dotted green curve shows the expected cumulative overlap if the modes were equally contributing each. The inset shows the initial structures of AMPAR (left) and NMDAR (right). Ionotropic glutamate receptors (iGluRs) mediate the majority of excitatory signal transmission in the brain. Their activation underlies learning and higher-order cognitive functions. iGluRs are organized in three domain layers: The trans-membrane domain (TMD) ion channel and two domains in the extracellular region (ECR) involved in ligand sensing - the ligand binding domain (LBD) and N-terminal domain (NTD). The NTD is a powerful allosteric module in NMDAR-type iGluRs but its function in the related AMPAR-type is less clear.
We performed structural dynamics analyses including all the three domains NTD-LBD-TMD, using the crystal structures of intact AMPAR (PDB: 3KG2) and NMDAR (PDB: 4PE5), based on elastic network models (Bahar et al. Annu. Rev. Biophys. 39, 23-42, 2010). We characterized the motions within and between domain layers, identifying movements that have been related to the gating properties and confirming a prominent motion underlying AMPAR desensitization using cysteine cross-linking in vivo. Our study disclosed conserved mechanisms of motions between the two receptors, despite their substantial differences in domain packing (RMSD ~ 18 �). The figure shows one of our computations for the correlation cosines between their structural difference and each of the softest 100 ANM modes accessible to the AMPAR in order to assess the ability of the two iGluR subtypes to interconvert conformations. It is noticeable that a single ANM mode 6 yields a correlation cosine of more than 0.50 - higher than that of a random mode by a factor of 40. These data show that AMPARs possess an intrinsic potential to 'easily' (via soft modes) convert to the NMDAR structure (that is, easy compression of the AMPAR towards the more tightly packed NMDAR). Moreover, we identified hotspot residues that are strategically localized to mediate allosteric signal transmission throughout the domain layers down to the channel gate. Overall, our data provided a first glimpse into the dynamic spectrum of AMPAR and NMDAR and delineated conserved mechanisms underlying allosteric communication in iGluRs.
Go to movies of global dynamics of AMPAR and NMDAR
Dutta A, Krieger J, Lee JY, Garcia-Nafria J, Greger IH, Bahar I. (2015) Cooperative Dynamics of Intact AMPA and NMDA Glutamate Receptors: Similarities and Subfamily-Specific Differences. Structure 23, 1692-1704. http://dx.doi.org/10.1016/j.str.2015.07.002
See more about MMBioS research in molecular modeling
Molecular Mechanism of Dopamine Transport by Human Dopamine Transporter
Dopamine transporter (DAT) controls neurotransmitter dopamine (DA) homeostasis by reuptake of excess DA from the synapse into the presynaptic neuron, assisted by the co-transport of two sodium ions. Malfunction of human DAT (hDAT) has been implicated in many neurological disorders. The first DAT structure (dDAT, from Drosophila melanogaster) has been recently resolved, which permit us to conduct a structure-based computational study of the time-resolved mechanism of DA reuptake by hDAT at full atomic scale. Using homology modeling and full-atomic microseconds accelerated simulation, we investigated the complete DA translocation including uptake from the EC region to its intracellular release and highlighted the key interactions that mediate DA translocation through hDAT. Our major observations are: spontaneous closure of extracellular gates prompted by DA binding; stabilization of a holo-occluded intermediate distinguished by close association of TM1b and TM6a with TM10 and resulting cluster of hydrophobic residues that prevents the hydration of the binding site; subsequent exposure to intracellular water triggered by Na2 dislocation, accompanied by a redistribution of salt bridges at the cytosolic surface; concerted tilting of TM5 and TM7, critical to opening the exit pore for substrate/ion channeling, enabled by the stretching of the G258-G263 loop, disruption of N82-N353 hydrogen bond, which drives the release of a Na+ ion and the Cl- ion; and DA release induced only after protonation of D79. Following Figure illustrates the transport mechanism, deduced from our over 13 sets micro-seconds MD simulations.
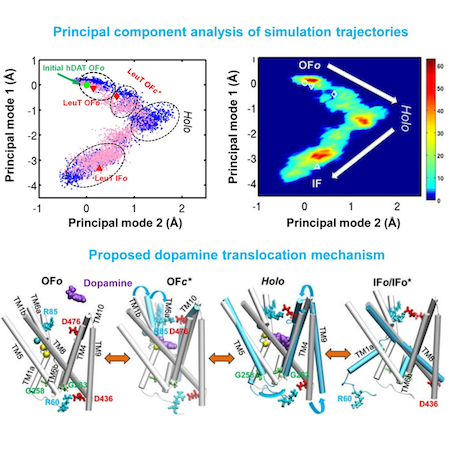
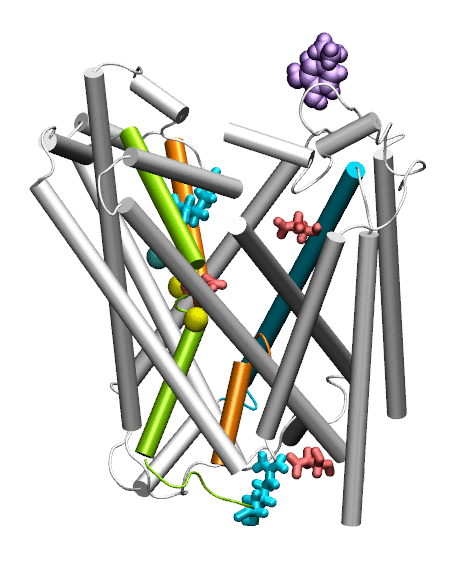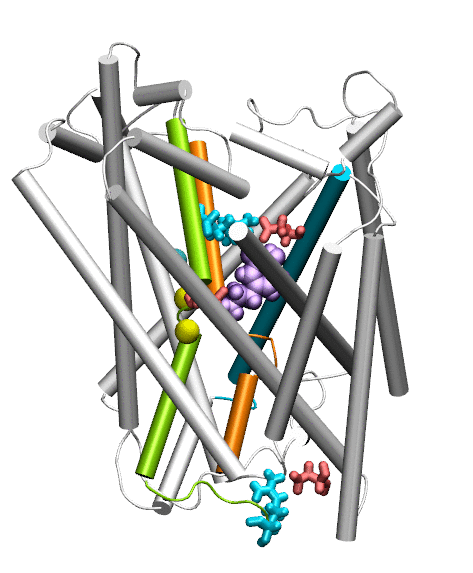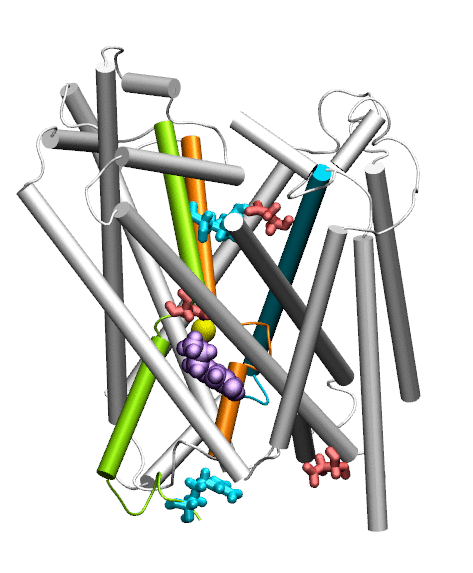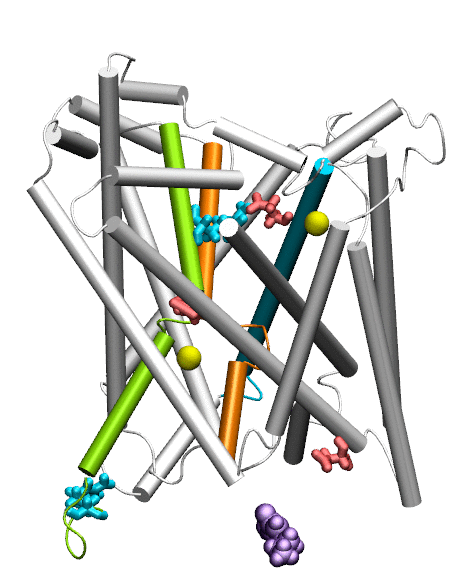
Movie: Dopamine extracellular binding, translocation, and intracellular release, captured by MD simlutions.
Cheng MH, Bahar I (2015) Molecular Mechanism of Dopamine Transport by Human Dopamine Transporter Structure 23: 2171-81.
See more about MMBioS research in molecular modeling
Short-Term Synaptic Facilitation Revealed
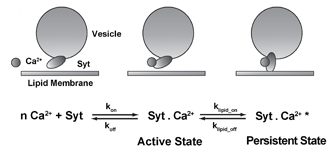
Short-term facilitation at synapses occurs during high-frequency stimulation, is known to be dependent on presynaptic calcium ions, and persists for tens of milliseconds after a presynaptic action potential. In a recent computational study, we have used the frog neuromuscular junction as a model synapse for both experimental and computer simulation studies aimed at testing various mechanistic hypotheses proposed to underlie short-term synaptic facilitation.
Our study built on our recently developed excess-calcium-binding-site model of synaptic vesicle release at the frog neuromuscular junction. Using MCell simulations, we investigated several mechanisms of short-term facilitation at the frog neuromuscular junction. Our studies place constraints on previously proposed facilitation mechanisms and conclude that the presence of a second class of calcium sensor proteins distinct from synaptotagmin can explain known properties of facilitation observed at the frog neuromuscular junction. We were further able to identify a novel facilitation mechanism, which relied on the persistent binding of calcium-bound synaptotagmin molecules to lipids of the presynaptic membrane. In a real physiological context, both mechanisms identified in our study - as well as others - may act simultaneously to give rise to the experimentally observed facilitation.
Thus, by combining computer simulations and physiological recordings we have developed a stochastic computer model of synaptic transmission at the frog neuromuscular junction, which enables microscopic insight into the biochemical dynamics of this model synapse and sheds light on its facilitation mechanism.
Ma J, Kelly L, Ingram J, Price TJ, Meriney SD, Dittrich M (2015) New insights into short-term synaptic facilitation at the frog neuromuscular junction Journal of Neurophysiology, 113:71-87.
See more about MMBioS research in cell modeling.
2014 Highlights
Learning Sequence Determinants of Protein:protein Interaction Specificity with Sparse Graphical Models
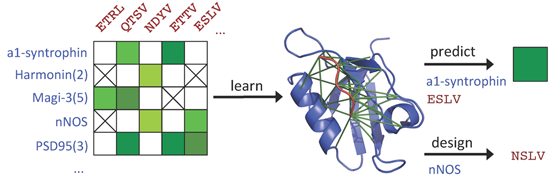 In studying the strength and specificity of interaction between members of two protein families, key questions center on which pairs of possible partners actually interact, how well they interact, and why they interact while others do not. Recently, we developed a method, DgSpi (Data-driven Graphical models of Specificity in Protein:protein Interactions), for learning and using graphical models that explicitly represent the amino acid basis for interaction specificity (why) and extend earlier classification-oriented approaches (which) to predict the free energy of binding (how well). Our predicted free energy values are highly predictive of the experimentally measured ones, reaching correlation coefficients of 0.69 in 10-fold cross-validation. Furthermore, the model serves as a compact representation of amino acid constraints underlying the interactions, enabling protein-level free energy predictions to be naturally understood in terms of residue-level constraints. Finally, DgSpi readily enables the design of new interacting partners, and we demonstrated that designed ligands are novel and diverse.
In studying the strength and specificity of interaction between members of two protein families, key questions center on which pairs of possible partners actually interact, how well they interact, and why they interact while others do not. Recently, we developed a method, DgSpi (Data-driven Graphical models of Specificity in Protein:protein Interactions), for learning and using graphical models that explicitly represent the amino acid basis for interaction specificity (why) and extend earlier classification-oriented approaches (which) to predict the free energy of binding (how well). Our predicted free energy values are highly predictive of the experimentally measured ones, reaching correlation coefficients of 0.69 in 10-fold cross-validation. Furthermore, the model serves as a compact representation of amino acid constraints underlying the interactions, enabling protein-level free energy predictions to be naturally understood in terms of residue-level constraints. Finally, DgSpi readily enables the design of new interacting partners, and we demonstrated that designed ligands are novel and diverse.
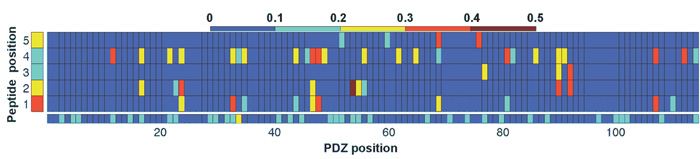
Kamisetty H, Ghosh B, Langmead CJ and Bailey-Kellogg C. Learning Sequence Determinants of Protein: Protein Interaction Specificity with Sparse Graphical Models. Proceedings of the 18th Annual International Conference Research in Computational Molecular Biology (RECOMB 2014) 129-143. PMCID: PMC4235964
See more about MMBioS research in molecular modeling
Advancing Parallel Bio-simulations
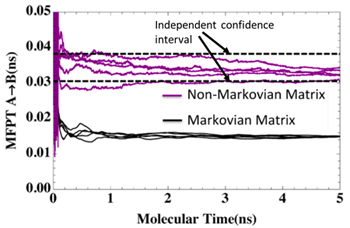
A central challenge in computer simulation is the extraction of timescales longer than the time for which biological systems can be simulated. The weighted ensemble (WE) parallel simulation approach, previously developed for a restricted set of problems, was recently advanced in MMBios work. Suarez and coworkers showed that a new non-Markovian analysis is capable of eliminating bias in estimates of long-timescale behavior, such as the mean first-passage time (MFPT) shown in the figure for the dissociation of methane molecules in explicit solvent. The analysis permits extraction of both equilibrium and non-equilibrium quantities.
Suarez E, Lettieri S, Zwier MC, Stringer CA, Subramanian SR, Chong LT, Zuckerman DM (2014) Simultaneous Computation of Dynamical and Equilibrium Information Using a Weighted Ensemble of Trajectories J. Chem. Theory Comput. 10:2658−2667. PMCID: PMC4168800
See more about MMBioS research in molecular modeling
Stochastic Modeling and Analysis of Apoptotic Events
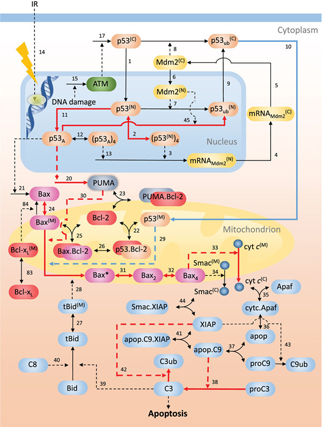
Developing pharmacological strategies for controlling ionizing radiation (IR)-induced cell death is important for both mitigating radiation damage and alleviating the side effects of anti-cancer radiotherapy manifested in surrounding tissue morbidity. Exposure to IR often triggers the onset of apoptotic events mediated by the tumor suppressor protein p53, which regulates IR-induced apoptosis via both transcription-dependent and -independent pathways. In a recent study, we constructed and calibrated a stochastic model of the p53 signaling network comprised of coupled modules of nuclear p53 activation, mitochondrial cytochrome c release and cytosolic caspase activation. The model takes into account cellular heterogeneity and is able to reproduce previously published experimental observations.
Through a detailed examination of the dynamics of p53 network in response to IR damage using stochastic simulation and statistical modeling checking techniques, we found that the strength of p53 transcriptional activity and its coupling (or timing with respect) to mitochondrial pore opening are major determinants of cell fate: for systems where apoptosis is elicited via a p53-transcription-independent mechanism, direct activation of Bax by p53 becomes critical to IR-induced-damage initiation. Our results also highlight the importance of a truncated Bid/caspase-3 feedback loop in driving apoptosis and determining treatment efficacy: if the feedback is sufficiently strong, inhibition of pro-apoptotic proteins like PUMA mitigates damage; but the effect weakens with delay in inhibitor administration. Further, we show that the combined inhibition of Bid and Bax elicits an anti-apoptotic response that is effective over a range of time delays. These results offer new insights into novel polypharmacological strategies for alleviating IR damage as well as controlling cell susceptibility to apoptosis under different disease states and environmental challenges.
Liu B, Bhatt D, Oltvai ZN, Greenberger JS, Bahar I (2014) Significance of p53 dynamics in regulating apoptosis in response to ionizing radiation, and polypharmacological strategies Sci. Rep. 4: 6245
See more about MMBioS research in cell modeling.
2013 Highlights
Gating events in LeuT
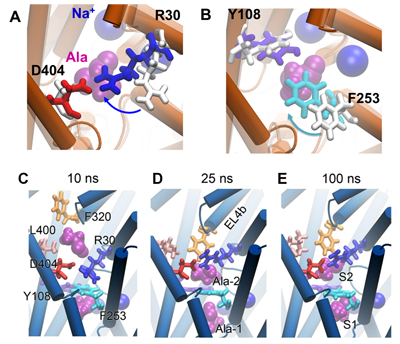
Top panel: Substrate Ala (purple vDW) binding to primary site S1 prompts the closure of EC gates. (A) Salt bridge formed intermittently between R30 (blue) and D404 (red); (B) Isomerization of F253 (cyan) brings its aromatic side chain into close proximity of Y108 (violet). White sticks show the side chain orientations in the OFo crystal structure. Results are from aMD simulation of Ala binding from the EC solution.
Bottom panel: The secondary substrate-binding site S2, the functional relevance of which has been debated, appears to stably bind an alanine only when the transporter assumes an intermediate conformer close to IF state. Results are from cMD simulations of IF state.
Unraveling the molecular mechanism of function of NSS family members has been a challenge due to the involvement of both local (EC or IC gate opening/closure) and global (between outward- and inward-facing) changes in structure. These events usually occur at different time scales (e.g. tens of nanoseconds for local, microseconds or slower for global). Their examination necessitates to adoption of multiscale methods. We implemented a multiscale approach that combines conventional molecular dynamics (MD), targeted MD and accelerated MD (aMD), to investigate substrate binding events from the extracellular region. Our study shed light at the same time at a controversial issue, the role of a secondary site for binding substrate, that have drawn much attention in recent years.
Mary H. Cheng and Bahar I (2013) Coupled Global and Local Changes Direct Substrate Translocation by Neurotransmitter-Sodium Symporter Ortholog LeuT Biophys J. 105:630-639. PMID: 23931311
Anton simulations confirm the global motions predicted by ANM for GltPh
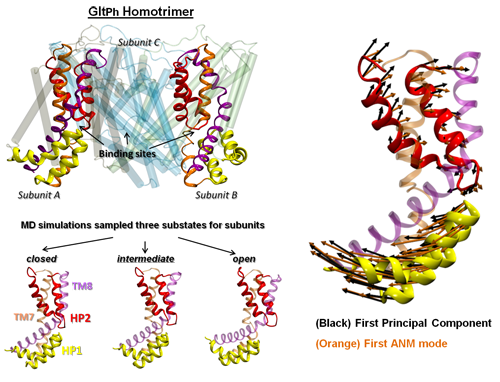 GltPh is the only structurally characterized member of the family of excitatory amino acid transporters. Even though various conformations of GltPh have been resolved to date and several simulations have been performed using these structural data, their mechanism of function is yet to be understood. GltPh is a homotrimer (left panel). Its monomers are composed each of two domains: a ‘scaffold’ (transmembrane TM helices 1, 2, 4 and 5) that provides support for the transport core and forms the interface between the monomers, and a transport core (TM3, 6, 7, 8 and two helical hairpins, HP1 and HP2), which binds and translocates the substrate and co-transported sodium ions. Substrate transport is enabled by alternating access to EC and IC environment, which is accomplished by the global transitions of GltPh between its outward-facing (OF) and inward-facing (IF) states, during substrate uptake and release, respectively. Experimental and computational studies have pointed to the role of HP1 and HP2 in gating the substrate-binding site. The conformations of HP1 and HP2 (bottom left panel) are determinants of the functional substates sampled during two Anton trajectories of 6 µs each generated for the IF state of GltPh. The essential motions extracted from the Anton simulations for GltPh in the IF state show close overlap with low frequency modes of Anisotropic Network Model (ANM) (right panel), a simple physics-based model of beads and springs, which exclusively depends on inter-residue contact topology.
GltPh is the only structurally characterized member of the family of excitatory amino acid transporters. Even though various conformations of GltPh have been resolved to date and several simulations have been performed using these structural data, their mechanism of function is yet to be understood. GltPh is a homotrimer (left panel). Its monomers are composed each of two domains: a ‘scaffold’ (transmembrane TM helices 1, 2, 4 and 5) that provides support for the transport core and forms the interface between the monomers, and a transport core (TM3, 6, 7, 8 and two helical hairpins, HP1 and HP2), which binds and translocates the substrate and co-transported sodium ions. Substrate transport is enabled by alternating access to EC and IC environment, which is accomplished by the global transitions of GltPh between its outward-facing (OF) and inward-facing (IF) states, during substrate uptake and release, respectively. Experimental and computational studies have pointed to the role of HP1 and HP2 in gating the substrate-binding site. The conformations of HP1 and HP2 (bottom left panel) are determinants of the functional substates sampled during two Anton trajectories of 6 µs each generated for the IF state of GltPh. The essential motions extracted from the Anton simulations for GltPh in the IF state show close overlap with low frequency modes of Anisotropic Network Model (ANM) (right panel), a simple physics-based model of beads and springs, which exclusively depends on inter-residue contact topology.
M Gur, Zomot E, and Bahar I (2013) Global Motions Exhibited by Proteins in Micro- to Milliseconds Simulations Concur with Anisotropic Network Model. J Chem Phys 139 (12):121912. PMCID: PMC3739829
On the Intracellular Gating of Glutamate Transporters
Sodium-coupled neurotransmitter transporters play a key role in neuronal signaling by clearing excess transmitter from the synapse. Structural data on a trimeric archaeal aspartate transporter, GltPh, have provided valuable insights into structural features of human excitatory amino acid transporters. However, the time-resolved mechanisms of substrate binding and release, as well as that of coupling to sodium co-transport, remain largely unknown for this important family.
Our recent study highlights the role of the helical hairpin HP2 as an intracellular gate, in addition to its role as an extracellular gate. In a recent study, we have elucidated the pathway of aspartate release from the inward-facing structure to the intracellular medium by generating multiple microsecond trajectories (using the Anton supercomputer), which consistently show that the helical hairpin HP2, not HP1, serves as an intracellular gate (in addition to its extracellular gating role). In contrast to previous proposals, HP1 can neither initiate nor accommodate neurotransmitter release without prior opening of HP2 by at least 4.0 Å. Aspartate release invariably follows that of a sodium ion located near the HP2 gate entrance. Asp394 on TM8 and Arg276 on HP1 emerge as key residues that promote the reorientation and diffusion of substrate toward the cell interior.
Zomot E and Bahar I (2013) Intracellular Gating in an Inward-facing State of Aspartate Transporter GltPh Is Regulated by the Movements of the Helical Hairpin HP2 J. Biol. Chem. 288:8231-37
See more about MMBioS research in molecular modeling and in modeling neurotransmitter transport.
Junction Crossing Ahead
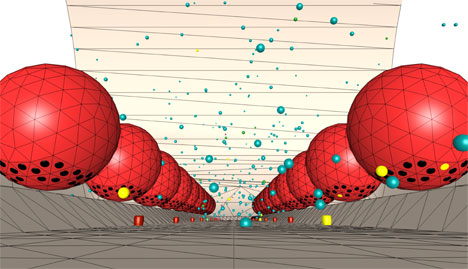 Synapses are the connections between neurons in the brain or between neurons and muscle fibers - so called neuromuscular junctions - and thus play a crucial role in human health. Despite decades of intense experimental study we still lack a detailed understanding of how synapses work.
Synapses are the connections between neurons in the brain or between neurons and muscle fibers - so called neuromuscular junctions - and thus play a crucial role in human health. Despite decades of intense experimental study we still lack a detailed understanding of how synapses work.
Many neuro-degenerative diseases such as dementia, depression and anxiety can be traced to improperly functioning synapses. Using a combination of computer simulations and experimental study, we have recently developed a spatially-realistic model of an active zone in the neuromuscular junction of the frog [1]. Using MCell's Monte Carlo algorithms to simulate the model, we could predict the calcium binding stoichiometry and dynamics that underlie
neurotransmitter release. Our model revealed that 20-40 independent calcium binding sites on synaptic vesicles, only a fraction of which need to bind calcium to trigger fusion, are sufficient to predict physiological release.
Our excess-calcium-binding site model has many functional advantages, agrees with recent data on synaptotagmin copy number, and is the first to link detailed physiological observations with the molecular machinery of calcium-triggered exocytosis. Importantly, our model provides detailed microscopic insight into the underlying calcium dynamics during synapse activation and thus constitutes a stepping stone for more detailed investigations in the future.
Dittrich M, Pattillo JM, King JD, Cho S, Stiles JR , Meriney SD (2013) An Excess-Calcium-Binding-Site Model Predicts Neurotransmitter Release at the Neuromuscular Junction Biophysical Journal 104:2751-2763.
See more about MMBioS research in cell modeling.
Stochastic Simulation
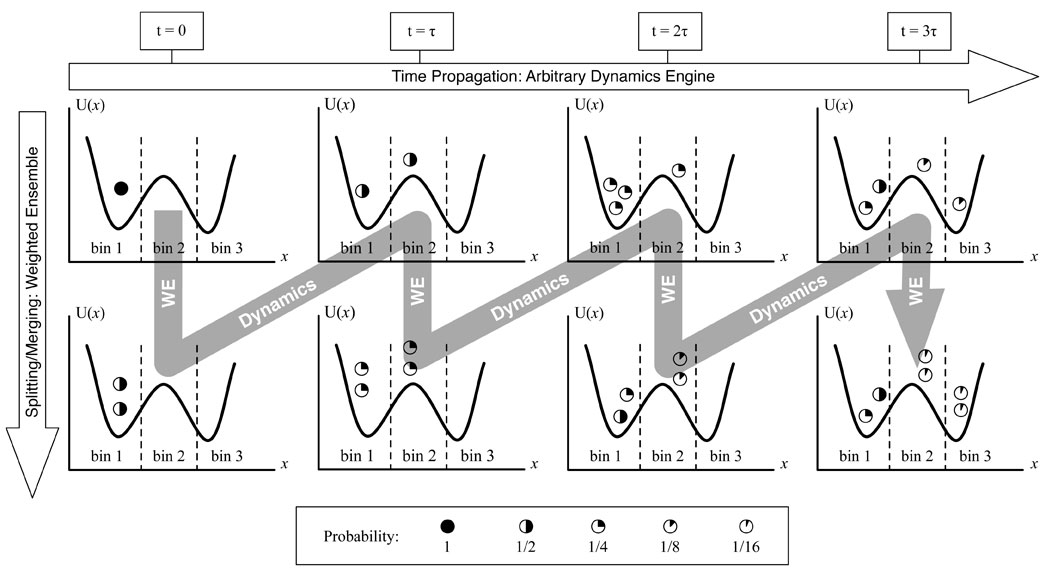 Systems biology, the quantitative study of complex interacting biological systems, is becoming increasingly demanding of computational resources. As models become able to capture truly physiological behavior, some of the most difficult computations may also be the most important, such as a rare transition from normal to pathological behavior.
Systems biology, the quantitative study of complex interacting biological systems, is becoming increasingly demanding of computational resources. As models become able to capture truly physiological behavior, some of the most difficult computations may also be the most important, such as a rare transition from normal to pathological behavior.
A novel approach to meet growing computational demands is described in a paper recently accepted to the Journal of Chemical Physics ("Efficient Stochastic Simulation of Chemical Kinetics Networks Using A Weighted Ensemble Of Trajectories"). First-author Donovan, in collaboration with MMBioS investigators Faeder and Zuckerman, applied a sophisticated parallel simulation algorithm originally developed for small-scale molecular systems to systems biology models.
The results were eye-catching: the new approach was more computationally efficient at modeling rare events than standard methods by orders of magnitude, even for a very complex model with thousands of reacting species.
Donovan RM, Sedgewick AJ, Faeder JR, Zuckerman DM (2013) Efficient Stochastic Simulation of Chemical Kinetics Networks Using A Weighted Ensemble Of Trajectories J. Chem. Phys. 139:115105.
See more about MMBioS research in molecular modeling.
CoMD: a Hybrid Methodology for Allosteric Changes
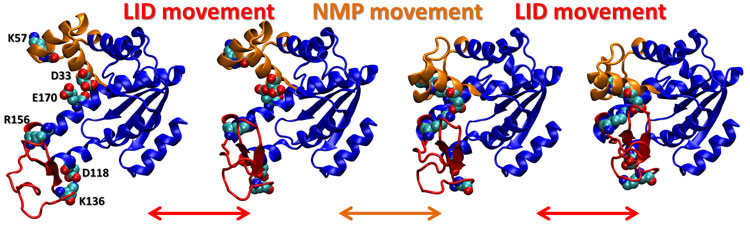
(Above) Three step (LID-NMP-LID) transition mechanism of adenylate kinase. Residues whose interactions undergo notable redistributions are shown in space-filling and labeled.
Efficient and accurate mapping of transition pathways is a challenging problem in allosteric proteins. We propose here a new methodology, called collective molecular dynamics (coMD) that takes advantage of the collective modes of motions encoded by the fold, while evaluating the interactions and energetics via a full-atomic molecular dynamics simulation protocol. The basic approach of the coMD is to deform the structure collectively along the modes predicted by the anisotropic network model (ANM)1 upon selecting them via a Monte Carlo/Metropolis algorithm from amongst the complete pool of all accessible modes. The method consists of a series of cycles, denoted by index k. Pairs of intermediate conformers are simultaneously generated starting from both ends at each cycle k. Each cycle is composed of multiple collectivemoves, s = 0, 1, 2,…, stot(k) where all Ca atoms are simultaneously displaced, guided by the ANM. The Ca-coordinates of the stot(k)th conformer are used to guide the full-atomic targeted molecular dynamics simulations (TMD), which followed by as short energy minimization (EM).
Application to adenylate kinase (AK), an allosteric enzyme composed of three domains, CORE, LID and NMP, shows that both open-to-closed and closed-to-open transitions of AK are readily sampled by coMD, being dominated by large-scale motions of the LID. An energy-barrier crossing occurs during the NMP movements. The energy barrier originates from a switch between the salt bridges K136- D118 at LID-CORE interface and K57-E170 and D33-R156 at CORE-NMP and LID-NMP interfaces, respectively. Despite its simplicity and computing efficiency, coMD yields ensembles of transition pathways in close accord with detailed full atomic simulations, lending support to its utility as a multiscale hybrid method for efficiently exploring the allosteric transitions of multidomain or multimeric proteins.
See more about MMBioS research in molecular modeling.
Micro-to-milliseconds Simulations vs Elastic Network Models
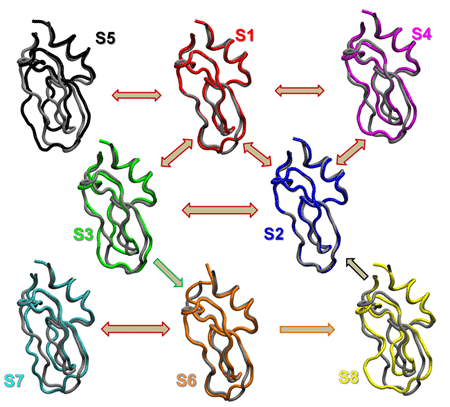
(Above) Structures representative of substates S1-S8 (colored) superimposed onto the crystal structure (grey). Transitions observed during simulations are indicated by arrows.
The Anton supercomputing technology recently developed for efficient molecular dynamics (MD) simulations permits us to examine micro- to millisecond events at full atomic resolution for proteins in explicit water and lipid bilayer. It also permits us to investigate to what extent the collective motions predicted by network models (that have found broad use in molecular biophysics) agree with those exhibited by full-atomic long simulations. The present study focuses on Anton trajectories generated for two systems: the bovine pancreatic trypsin inhibitor (BPTI), and an archaeal aspartate transporter, GltPh. The former, a thoroughly studied system, helps benchmark the method of comparative analysis, and the latter provides new insights into the mechanism of function of glutamate transporters. The principal components (PCs) of motion derived from the 1.013 ms long MD trajectory of BPTI closely overlapped with those predicted by the anisotropic network model (ANM)1, which a simple physics-based model of beads and springs that exclusively depends on inter-residue contact topology.
Notably, the ANM modes define the collective mechanisms, or the pathways on conformational energy landscape, that underlie the passage between the crystal structure and substates visited in simulations. In particular, the lowest frequency ANM modes facilitate the conversion between the most probable substates, lending support to the view that easy access to functional substates is a robust determinant of evolutionarily selected native contact topology.
Similar to BPTI, the PCs of the 12 µs long MD trajectory of the GltPh homotrimer3 exhibited good overlap with its ANM modes. Moreover, these ANM modes appear to naturally facilitate the transitions between the substates sampled in the simulations.
Gur M, Zomot E, Bahar I (2013) Global motions exhibited by proteins in micro- to milliseconds simulations concur with anisotropic network model predictions J Chem Phys 139(12): 121912.
See more about MMBioS research in molecular modeling.
ANMPathway for Exploring Transitions
Biomolecular conformational transitions are essential to biological functions. Most experimental methods report on the long-lived functional states of biomolecules, but information about the transition pathways between these stable states is generally scarce. Such transitions involve short-lived conformational states that are difficult to detect experimentally.
For this reason, computational methods are needed to produce plausible hypothetical transition pathways that can then be probed experimentally. Here we propose a simple and computationally efficient method, called ANMPathway, for constructing a physically reasonable pathway between two endpoints of a conformational transition. We adopt a coarse-grained representation of the protein and construct a two-state potential by combining two anisotropic network models (ANMs)1 representative of the experimental structures resolved for the endpoints.
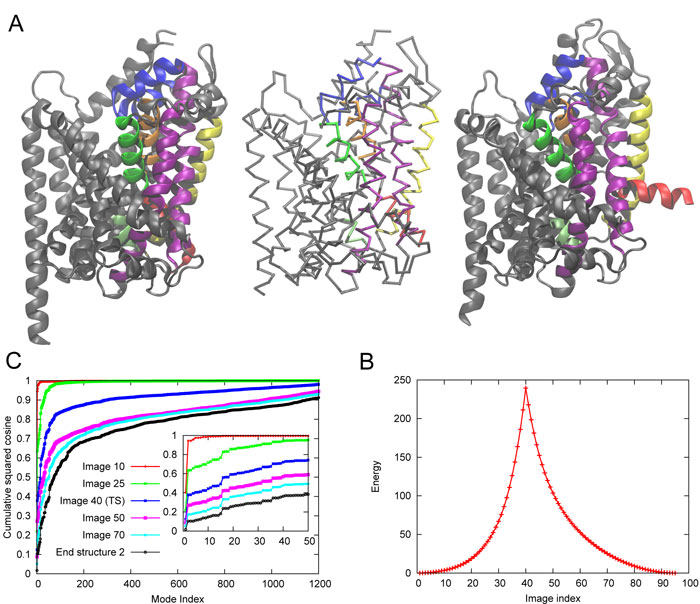
(Above) Conformational transition between states with two protomers facing inward and all protomers facing inward of the glutamate transporter, GltPh. (A). Left : Structure of the intermediate state (iOF) state where two protomers are in IF and the third (green) protomer is in the OF; right : IF state where all protomers are in IF. The central diagram (stick representation) is the Cα trace of the transition state produced by ANMPathway. (B) The energy of the system along the transition. Total number of images in the pathway is 79, RMSD between two consecutive images is ~0.1Å. (C) Cumulative squared cosines between the structural change to reach a few selected conformers/images along the transition pathway and the ANM modes accessible to the starting (iOF) structure. The force constants and cut-offs for both the end states were set to 0.1 kcal/mol and 15 Å respectively. No energy offsets were used for either of the end structures.
The two-state potential has a cusp in the configuration space where the energies from both the ENMs are equal. We first search for the minimum energy structure along the cusp hyper-surface and then treat it as the transition state. The continuous pathway is subsequently constructed by following the steepest descent energy minimization trajectories starting from the transition state on each side of the cusp. Application to several systems of broad biological interest such as adenylate kinase, ATP-driven calcium pump SERCA, leucine transporter and glutamate transporter (See figure at left) shows that ANMPathway yields results in good agreement with those from other similar methods and with data obtained from all-atom molecular dynamics simulations, in support of the utility of this simple and efficient approach. Notably the method provides experimentally testable predictions, including the formation of non-native contacts during the transition which we were able to detect in two of the systems we studied.
See more about MMBioS research in molecular modeling.
Major New Release of CellOrganizer 2.0 Published

(Above) Example cell image generated by CellOrganizer showing the nuclear membrane (red), cell boundary (blue) and individual lysosomes (yellow).
A major new release of the CellOrganizer system for creating image-derived models of cell shape and organization has just been published. The software is a major focus of research supported by the NIH through the National Center for Multiscale Modeling of Biological Systems (MMBioS). It creates statistical, generative models of cell and nuclear shape, microtubule patterns, and vesicular organelles that can be used as the basis for cell simulations (another focus of MMBioS). Generative models capture not just a description of a pattern but the ability to produce new examples of that pattern (analogously to the way in which humans capture models of letters or spoken words not just by describing them but by learning to produce new examples). Support for CellOrganizer has also come from NIH grants GM075205 and GM090033. Specific improvements or additions in the new release include:
Cell image synthesis
- Improved vesicular organelles model
- Eliminate object/object and object/boundary overlap during generation
- Ability to combine models learned from images of different resolution, and synthesize images at desired resolution
- Ability to synthesis random walks in shape space from diffeomorphic models of cell and nuclear shape
- Includes directed random walks using Willmore energy and shape space density
Model training
- New capabilities for cell and nuclear shape model learning
- Build nuclear models from images without nuclear marker
- Build joint diffeomorphic models of cell and nuclear shape
- Per-cell representations for easy model building and comparison
Other
- Export to SBML-spatial and mesh formats for interfacing with tools such as CellBlender and VCell
- Parallelization of model learning pipeline
- Reporter tools
- Assess learned models
- Compare models
- Compare per-cell parameters within or across models
See more about MMBioS research in image processing and analysis.
Release of AlignTK Image Alignment Software
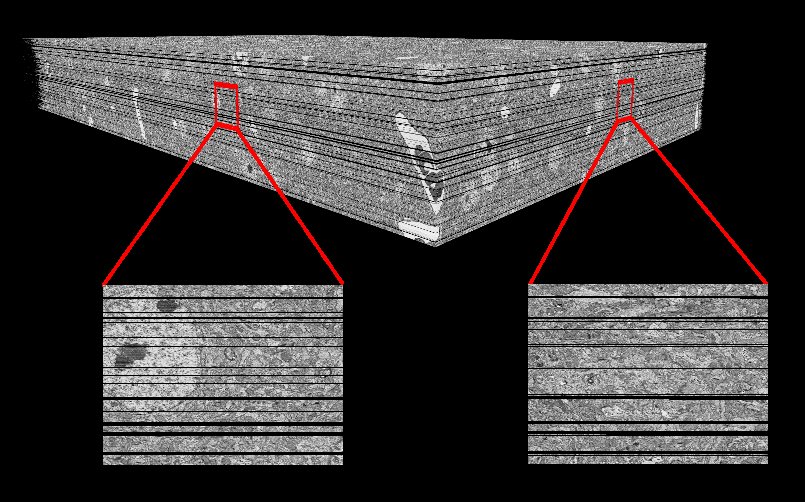

A cutaway view of a 450x350x52μm volume reconstructed from ~1150 45nm sections of mouse visual cortex. (Missing sections are left black.) Closeups of the x-z and y-z faces show that neural features can easily be traced in a direction perpendicular to the original cutting planes.
We have issued the first public release of the AlignTK toolkit for image alignment. This package is primarily targeted at large 3-D datasets that are acquired in sections, such as in serial-section electron microscopy. The sectioning process physically distorts each tissue section in a slightly different way, and imaging these sections typically introduces additional electron-beam and optical distortions. Aligning these sections to produce a coherent 3-D volume is challenging, and AlignTK provides a set of tools that can be used in a batch environment to process millions of camera images into a volumetric dataset in which axons and dendrites of individual neurons can be traced.
An earlier version of these tools was used at NRBSC and the Pittsburgh Supercomputing Center to align the mouse visual cortex dataset in the Nature paper by Bock et al. These tools have also been applied to optical images of stained mouse brain sections and time-series fluorescence images of the frog neuromuscular junction.
Bock, DD, Lee, WA, Kerlin, AM, Andermann, ML, Hood, G, Wetzel, AW, Yurgenson, S, Soucy, ER, Kim, HS, & Reid, RC (2011). Network anatomy and in vivo physiology of visual cortical neurons, Nature, 471:177-182.
See more about MMBioS research in image processing and analysis.